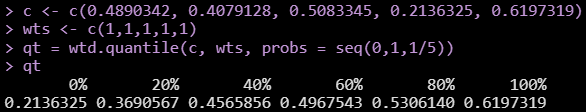
I want to convert the R package Hmisc::wtd.quantile() into python.
I took this as reference and it seems that the logics are different than R:
# First function
def weighted_quantile(values, quantiles, sample_weight = None,
values_sorted = False, old_style = False):
""" Very close to numpy.percentile, but supports weights.
NOTE: quantiles should be in [0, 1]!
:param values: numpy.array with data
:param quantiles: array-like with many quantiles needed
:param sample_weight: array-like of the same length as `array`
:return: numpy.array with computed quantiles.
"""
values = np.array(values)
quantiles = np.array(quantiles)
if sample_weight is None:
sample_weight = np.ones(len(values))
sample_weight = np.array(sample_weight)
assert np.all(quantiles >= 0) and np.all(quantiles <= 1), 'quantiles should be in [0, 1]'
if not values_sorted:
sorter = np.argsort(values)
values = values[sorter]
sample_weight = sample_weight[sorter]
# weighted_quantiles = np.cumsum(sample_weight)
# weighted_quantiles /= np.sum(sample_weight)
weighted_quantiles = np.cumsum(sample_weight)/np.sum(sample_weight)
return np.interp(quantiles, weighted_quantiles, values)
weighted_quantile(values = [0.4890342, 0.4079128, 0.5083345, 0.2136325, 0.6197319],
quantiles = np.arange(0, 1 1 / 5, 1 / 5),
sample_weight = [1,1,1,1,1])
>> array([0.2136325, 0.2136325, 0.4079128, 0.4890342, 0.5083345, 0.6197319])
# Second function
def weighted_percentile(data, weights, perc):
"""
perc : percentile in [0-1]!
"""
data = np.array(data)
weights = np.array(weights)
ix = np.argsort(data)
data = data[ix] # sort data
weights = weights[ix] # sort weights
cdf = (np.cumsum(weights) - 0.5 * weights) / np.sum(weights) # 'like' a CDF function
return np.interp(perc, cdf, data)
weighted_percentile([0.4890342, 0.4079128, 0.5083345, 0.2136325, 0.6197319], [1,1,1,1,1], np.arange(0, 1 1 / 5, 1 / 5))
>> array([0.2136325 , 0.31077265, 0.4484735 , 0.49868435, 0.5640332 ,
0.6197319 ])
Both are different with R. Any idea?
CodePudding user response:
I am Python-illiterate, but from what I see and after some quick checks I can tell you the following.
Here you use uniform (sampling) weights, so you could also directly use the quantile() function. Not surprisingly, it gives the same results as wtd.quantile() with uniform weights:
x <- c(0.4890342, 0.4079128, 0.5083345, 0.2136325, 0.6197319)
n <- length(x)
x <- sort(x)
quantile(x, probs = seq(0,1,0.2))
# 0% 20% 40% 60% 80% 100%
# 0.2136325 0.3690567 0.4565856 0.4967543 0.5306140 0.6197319
The R quantile() function get the quantiles in a 'textbook' way, i.e. by determining the index i of the obs to use with i = q(n 1).
In your case:
seq(0,1,0.2)*(n 1)
# 0.0 1.2 2.4 3.6 4.8 6.0
Of course since you have 5 values/obs and you want quintiles, the indices are not integers. But you know for example that the first quintile (i = 1.2) lies between obs 1 and obs 2. More precisely, it is a linear combination of the two observations (the 'weights' are derived from the value of the index):
0.2*x[1] 0.8*x[2]
# 0.3690567
You can do the same for the all the quintiles, on the basis of the indices:
q <-
c(min(x), ## 0: actually, the first obs
0.2*x[1] 0.8*x[2], ## 1.2: quintile lies between obs 1 and 2
0.4*x[2] 0.6*x[3], ## 2.4: quintile lies between obs 2 and 3
0.6*x[3] 0.4*x[4], ## 3.6: quintile lies between obs 3 and 4
0.8*x[4] 0.2*x[5], ## 4.8: quintile lies between obs 4 and 5
max(x) ## 6: actually, the last obs
)
q
# 0.2136325 0.3690567 0.4565856 0.4967543 0.5306140 0.6197319
You can see that you get exactly the output of quantile() and wtd.quantile().
If instead of 0.2*x[1] 0.8*x[2] we consider the following:
0.5*x[1] 0.5*x[2]
# 0.3107726
We get the output of your second Python function. It appears that your second function considers uniform 'weights' (obviously I am not talking about the sampling weights here) when combining the two observations. The issue (at least for the second Python function) seems to come from this. I know these are just insights, but I hope they will help.
EDIT: note that the difference between the two is not necessary an 'issue' with the python code. There are different quantile estimators (and their weighted versions) and the python functions could simply rely on a different estimator than Hmisc::wtd.quantile(). I think that the latter uses the weighted version of the Harrell-Davis quantile estimator. If you really want to implement this one, you should check the source code of Hmisc::wtd.quantile() and try to 'directly' translate this into Python.