Hi I'm analysing the pattern of spending for individuals before they died. My dataset contains individuals' monthly spending and their dates of death. The dataset looks similar to this:
ID 2018_11 2018_12 2019_01 2019_02 2019_03 2019_04 2019_05 2019_06 2019_07 2019_08 2019_09 2019_10 2019_11 2019_12 2020_01 date_of_death
A 15 14 6 23 23 5 6 30 1 15 6 7 8 30 1 2020-01-02
B 2 5 6 7 7 8 9 15 12 14 31 30 31 0 0 2019-11-15
Each column denotes the month of the year. For example, "2018_11" means November 2018. The number in each cell denotes the spending in that specific month.
I would like to construct a data frame which contains the spending data of each individual in their last 0-12 months. It will look like this:
ID last_12_month last_11_month ...... last_1_month last_0_month date_of_death
A 6 23 30 1 2020-01-02
B 2 5 30 31 2019-11-15
Each individual died at different time. For example, individual A died on 2020-01-02, so the data of the "last_0_month" for this person should be extracted from the column "2020_01", and that of "last_12_month" extracted from "2019_01"; individual B died on 2019-11-15, so the data of "last_0_month" for this person should be extracted from the column "2019_11", and that of "last_12_month" should be extracted from the column "2018_11".
I will be really grateful for your help.
CodePudding user response:
Here is a tidyverse solution.
Reshape the data to long format, coerce the date columns to class "Date", use Dirk Eddelbuettel's accepted answer to this question to compute the date differences in months and keep the rows with month differences between 0 and 12.
This grouped long format is probably more useful and I compute means by group and plot the spending of the last 12 months prior to death but since the question asks for a wide format, the output data set spending12_wide is created.
options(width=205)
df1 <- read.table(text = "
ID 2018_11 2018_12 2019_01 2019_02 2019_03 2019_04 2019_05 2019_06 2019_07 2019_08 2019_09 2019_10 2019_11 2019_12 2020_01 date_of_death
A 15 14 6 23 23 5 6 30 1 15 6 7 8 30 1 2020-01-02
B 2 5 6 7 7 8 9 15 12 14 31 30 31 0 0 2019-11-15
", header = TRUE, check.names = FALSE)
suppressPackageStartupMessages(library(dplyr))
library(tidyr)
library(ggplot2)
# Dirk's functions
monnb <- function(d) {
lt <- as.POSIXlt(as.Date(d, origin = "1900-01-01"))
lt$year*12 lt$mon
}
# compute a month difference as a difference between two monnb's
diffmon <- function(d1, d2) { monnb(d2) - monnb(d1) }
spending12 <- df1 %>%
pivot_longer(cols = starts_with('20'), names_to = "month") %>%
mutate(month = as.Date(paste0(month, "_01"), "%Y_%m_%d"),
date_of_death = as.Date(date_of_death)) %>%
group_by(ID, date_of_death) %>%
mutate(diffm = diffmon(month, date_of_death)) %>%
filter(diffm >= 0 & diffm <= 12)
spending12 %>% summarise(spending = mean(value), .groups = "drop")
#> # A tibble: 2 x 3
#> ID date_of_death spending
#> <chr> <date> <dbl>
#> 1 A 2020-01-02 12.4
#> 2 B 2019-11-15 13.6
spending12_wide <- spending12 %>%
mutate(month = zoo::as.yearmon(month)) %>%
pivot_wider(
id_cols = c(ID, date_of_death),
names_from = diffm,
names_glue = "last_{.name}_month",
values_from = value
)
spending12_wide
#> # A tibble: 2 x 15
#> # Groups: ID, date_of_death [2]
#> ID date_of_death last_12_month last_11_month last_10_month last_9_month last_8_month last_7_month last_6_month last_5_month last_4_month last_3_month last_2_month last_1_month last_0_month
#> <chr> <date> <int> <int> <int> <int> <int> <int> <int> <int> <int> <int> <int> <int> <int>
#> 1 A 2020-01-02 6 23 23 5 6 30 1 15 6 7 8 30 1
#> 2 B 2019-11-15 2 5 6 7 7 8 9 15 12 14 31 30 31
ggplot(spending12, aes(month, value, color = ID))
geom_line()
geom_point()
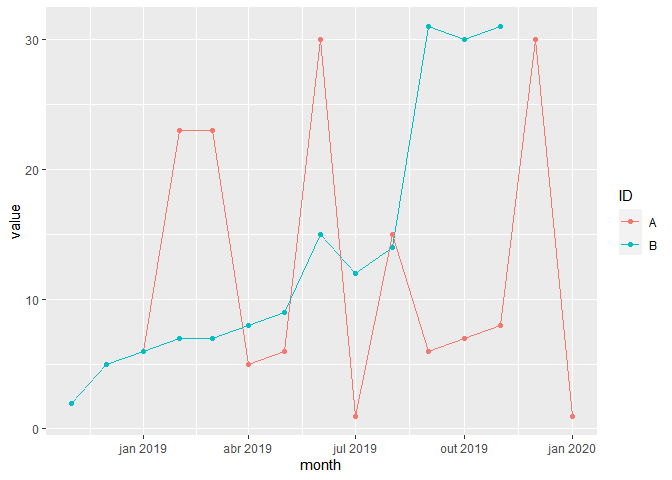
Created on 2022-03-09 by the reprex package (v2.0.1)
CodePudding user response:
here you can find a similar approach to the one presented by @RuiBarradas but using lubridate for extracting the difference in months:
library(dplyr)
library(tidyr)
library(lubridate)
# Initial data
df <- structure(list(
ID = c("A", "B"),
`2018_11` = c(15, 2),
`2018_12` = c(14, 5),
`2019_01` = c(6, 6),
`2019_02` = c(23, 7),
`2019_03` = c(23, 7),
`2019_04` = c(5, 8),
`2019_05` = c(6, 9),
`2019_06` = c(30, 15),
`2019_07` = c(1, 12),
`2019_08` = c(15, 14),
`2019_09` = c(6, 31),
`2019_10` = c(7, 30),
`2019_11` = c(8, 31),
`2019_12` = c(30, 0),
`2020_01` = c(1, 0),
date_of_death = c("2020-01-02", "2019-11-15")
),
row.names = c(NA, -2L),
class = "data.frame"
)
# Convert to longer all cols that start with 20 (e.g. 2020, 2021)
df_long <- df %>%
pivot_longer(starts_with("20"), names_to = "month")
# treatment
df_long <- df_long %>%
mutate(
# To date, just in case
date_of_death = as.Date(date_of_death),
# Need to reformat the colnames from (e.g.) 2021_01 to 2021-01-01
month_fmt = as.Date(paste0(gsub("_", "-", df_long$month), "-01")),
# End of month
month_fmt = ceiling_date(month_fmt, "month") - days(1),
# End of month for month of death
date_of_death_eom = ceiling_date(date_of_death, "month") - days(1),
# Difference in months (using end of months
month_diff = round(time_length(
interval(month_fmt, date_of_death_eom),"month"),0)) %>%
# Select only months bw 0 and 12
filter(month_diff %in% 0:12) %>%
# Create labels for the next step
mutate(labs = paste0("last_", month_diff,"_month"))
# To wider
end <- df_long %>%
pivot_wider(
id_cols = c(ID, date_of_death),
names_from = labs,
values_from = value
)
end
#> # A tibble: 2 x 15
#> ID date_of_death last_12_month last_11_month last_10_month last_9_month
#> <chr> <date> <dbl> <dbl> <dbl> <dbl>
#> 1 A 2020-01-02 6 23 23 5
#> 2 B 2019-11-15 2 5 6 7
#> # ... with 9 more variables: last_8_month <dbl>, last_7_month <dbl>,
#> # last_6_month <dbl>, last_5_month <dbl>, last_4_month <dbl>,
#> # last_3_month <dbl>, last_2_month <dbl>, last_1_month <dbl>,
#> # last_0_month <dbl>
Created on 2022-03-09 by the reprex package (v2.0.1)
CodePudding user response:
Using data.table and lubridate packages
library(data.table)
library(lubridate)
setDT(dt)
dt <- melt(dt, id.vars = c("ID", "date_of_death"))
dt[, since_death := interval(ym(variable), ymd(date_of_death)) %/% months(1)]
dt <- dcast(dt[since_death 