I am working on an assignment, which tasks me to generate a list of data, using the below code.
##Use the make_data function to generate 25 different datasets, with mu_1 being a vector
x <- seq(0, 3, len=25)
make_data <- function(a){
n = 1000
p = 0.5
mu_0 = 0
mu_1=a
sigma_0 = 1
sigma_1 = 1
y <- rbinom(n, 1, p)
f_0 <- rnorm(n, mu_0, sigma_0)
f_1 <- rnorm(n, mu_1, sigma_1)
x <- ifelse(y == 1, f_1, f_0)
test_index <- createDataPartition(y, times = 1, p = 0.5, list = FALSE)
list(train = data.frame(x = x, y = as.factor(y)) %>% slice(-test_index),
test = data.frame(x = x, y = as.factor(y)) %>% slice(test_index))
}
dat <- sapply(x,make_data)
The code looks good to go, and 'dat' appears to be a 25 column, 2 row table, each with its own data frame.
Now, each data frame within a cell has 2 columns.
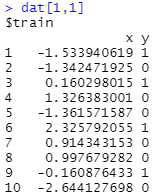
And this is where I get stuck.
While I can get to the data frame in row 1, column 1, just fine (i.e. just use dat[1,1]), I can't reach the column of 'x' values within dat[1,1]. I've experimented with
dat[1,1]$x
dat[1,1][1]
But they only throw weird responses: error/null.
Any idea how I can pull the column? Thanks.
CodePudding user response:
dat[1, 1] is a list.
class(dat[1, 1])
#[1] "list"
So to reach to x you can do
dat[1, 1]$train$x
Or
dat[1, 1][[1]]$x
As a sidenote, instead of having this 25 X 2 matrix as output in dat I would actually prefer to have a nested list.
dat <- lapply(x,make_data)
#Access `x` column of first list from `train` dataset.
dat[[1]]$train$x
However, this is quite subjective and you can chose whatever format you like the best.
