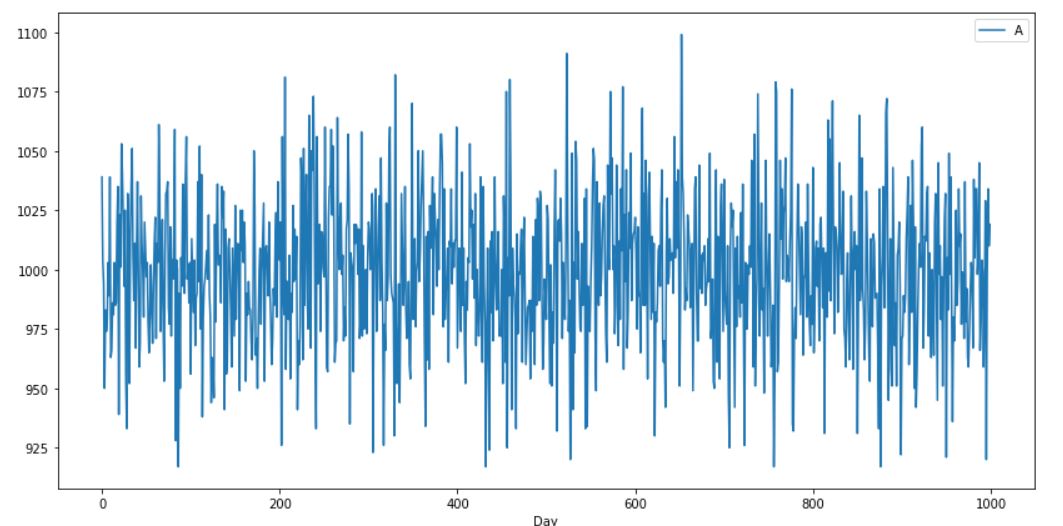
This is my model
# normalize the dataset
scaler = MinMaxScaler(feature_range=(0, 1))
dataset = scaler.fit_transform(df)
train_size = int(len(dataset) * 0.8)
test_size = len(dataset) - train_size
train = dataset[0:train_size,:]
test = dataset[train_size:len(dataset),:]
def create_dataset(dataset, look_back=1):
dataX, dataY = [], []
for i in range(len(dataset)-look_back-1):
a = dataset[i:(i look_back), 0]
dataX.append(a)
dataY.append(dataset[i look_back, 0])
return np.array(dataX), np.array(dataY)
# reshape into X=t and Y=t 1
look_back = 15
trainX, trainY = create_dataset(train, look_back)
testX, testY = create_dataset(test, look_back)
print(trainX.shape)
# reshape input to be [samples, time steps, features]
trainX = np.reshape(trainX, (trainX.shape[0], trainX.shape[1], 1))
testX = np.reshape(testX, (testX.shape[0], testX.shape[1], 1))
from keras.layers import Dropout
from keras.layers import Bidirectional
model=Sequential()
model.add(LSTM(50,activation='relu',return_sequences=True,input_shape=(look_back,1)))
model.add(LSTM(50, activation='relu', return_sequences=True))
model.add(LSTM(50, activation='relu', return_sequences=True))
model.add(LSTM(50, activation='sigmoid', return_sequences=False))
model.add(Dense(50))
model.add(Dense(50))
model.add(Dropout(0.2))
model.add(Dense(1))
model.compile(optimizer='adam',loss='mean_squared_error',metrics=['accuracy'])
model.optimizer.learning_rate = 0.0001
Xdata_train=[]
Ydata_train=[]
Xdata_train, Ydata_train = create_dataset(train, look_back)
Xdata_train = np.reshape(Xdata_train, (Xdata_train.shape[0], Xdata_train.shape[1], 1))
#training for all data
history = model.fit(Xdata_train,Ydata_train,batch_size=1,epochs=10,shuffle=False)
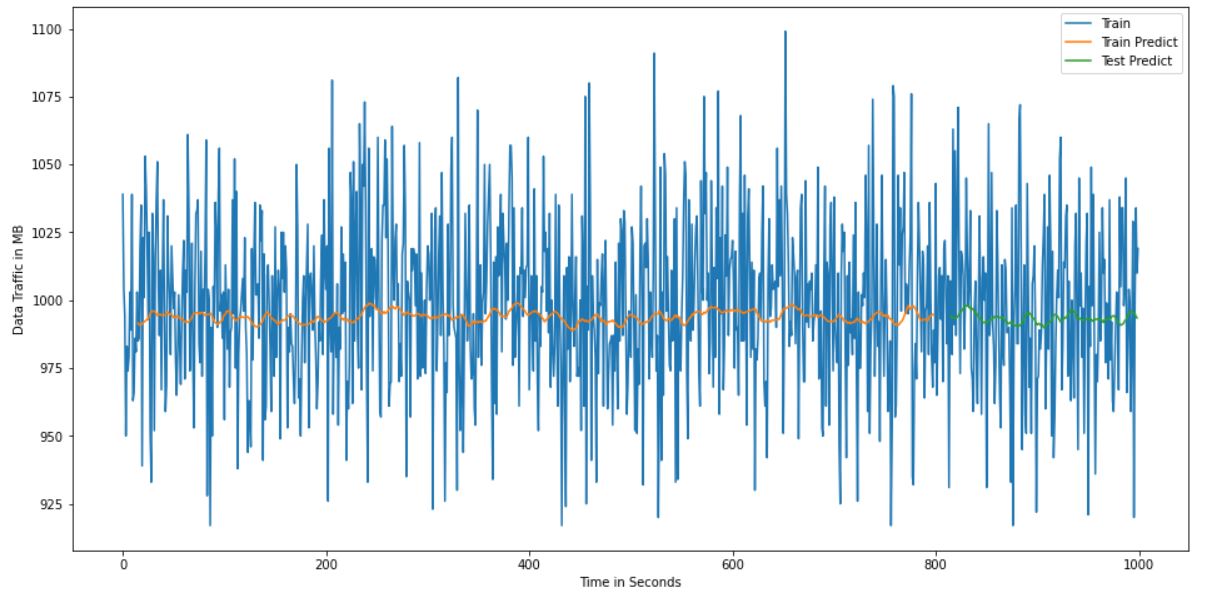
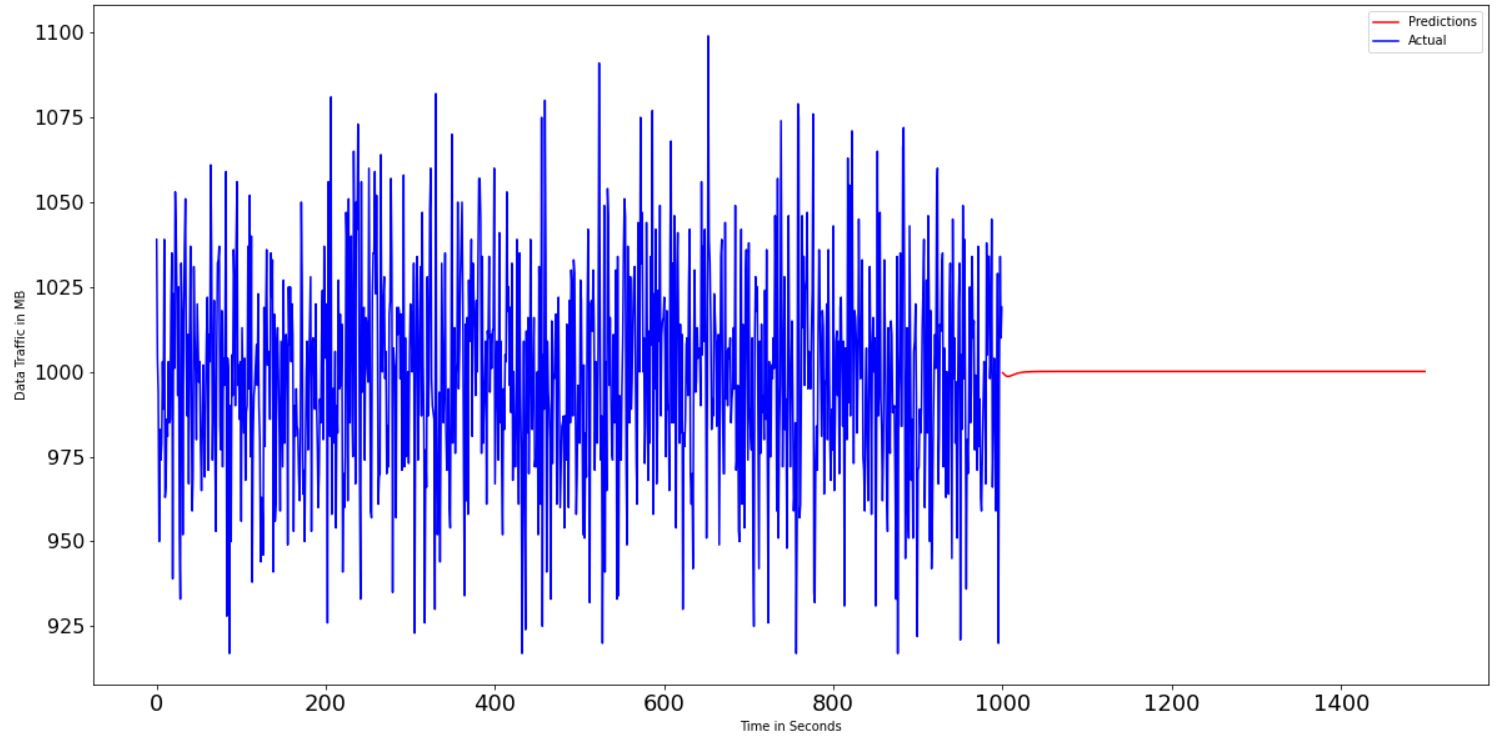
RMSE value is around 35 and accuracy is very low. When I icrese the epochs there is no any variation. What are the changes should I do to get the accuracy at high value. Here i attached the graphical results to get an idea.
How could I fix this?
CodePudding user response:
Just with a once-over on your code, I can think of a few of the changes. Try using Bidirectional LSTM, binary_cross_entropy (assuming it's a binary classification) for the loss, and shuffle = True on training. Also, try adding Dropout between LSTM layers.
CodePudding user response:
Here are couple of suggestions:
First of all, never fit normalizer on the entire dataset. First partition your data into train/test parts, fit the scaler on train data and then transform both train/test using that scaler. Otherwise you are leaking the information from your test data into training when doing the normalization (such as min/max values or std/mean when using standard scaler).
You seem to be normalizing your
ydata, but never reverting the normalization, as a result you end up with an output on lower scale (as we can see on plots). You can redo normalization usingscaler.inverse_transform().Finally, you may want to remove sigmoid activation function from the LSTM layer, its generally not a good idea to use sigmoid anywhere else besides the output layer as it may cause vanishing gradient.