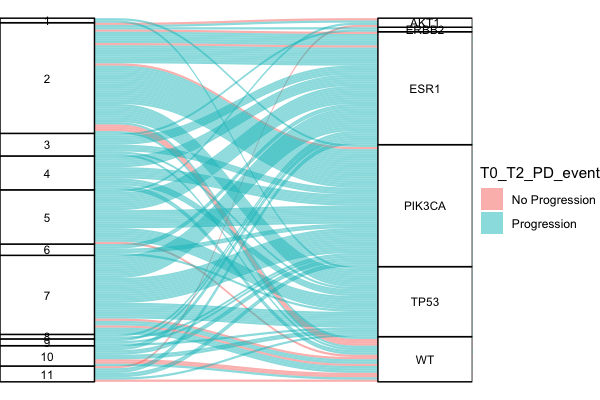
I am trying to figure out how can I change the scales of the columns box, making them bigger so I can see clearly the names.
Is also possible to paint the PIK3CA and ESR1 boxes in dark green and light orange, respectively? Thank you so much! Any help is more than welcome.
This is my code and the plot that I have generate:
library(ggalluvial)
ggplot(data = Allu,
aes(axis1 = Metastasis_Location, axis2 = Gene_mut, y = Freq))
geom_alluvium(aes(fill = T0_T2_PD_event),
curve_type = "quintic")
geom_stratum(width = 1/4)
geom_text(stat = "stratum", size = 3,
aes(label = after_stat(stratum)))
scale_x_discrete(limits = c("Metastasis_Location", "Gene_mut"),
expand = c(0.05, .05))
theme_void()
My data:
structure(list(Metastasis_Location = c(1L, 1L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 3L, 3L, 3L,
3L, 3L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L,
4L, 4L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L,
5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 6L, 6L,
6L, 6L, 6L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L,
7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L,
7L, 7L, 7L, 7L, 7L, 7L, 8L, 8L, 9L, 9L, 9L, 10L, 10L, 10L, 10L,
10L, 10L, 10L, 10L, 10L, 11L, 11L, 11L, 11L, 11L, 11L, 11L),
T0_T2_THERAPY_COD = structure(c(2L, 2L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L,
1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 1L, 1L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 1L, 1L, 2L, 2L, 2L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 1L, 2L, 2L,
2L, 2L, 2L, 2L), .Label = c("A", "F"), class = "factor"),
T0_T2_PD_event = structure(c(2L, 2L, 1L, 1L, 1L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 2L, 2L, 2L, 2L, 2L, 2L,
1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 1L, 2L, 2L, 2L, 1L, 2L, 2L, 2L, 2L, 1L, 1L, 2L,
2L, 2L, 2L, 2L), .Label = c("No Progression", "Progression"
), class = "factor"), Gene_mut = structure(c(4L, 5L, 1L,
3L, 4L, 1L, 2L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 4L, 4L,
4L, 5L, 5L, 5L, 6L, 3L, 6L, 6L, 6L, 3L, 3L, 3L, 3L, 3L, 3L,
3L, 3L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 5L, 5L, 5L,
5L, 5L, 6L, 2L, 3L, 4L, 4L, 3L, 3L, 3L, 4L, 5L, 6L, 3L, 6L,
3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 6L, 3L, 4L,
4L, 5L, 6L, 1L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L,
4L, 4L, 5L, 5L, 5L, 5L, 5L, 3L, 4L, 3L, 4L, 5L, 6L, 3L, 3L,
4L, 5L, 6L, 6L, 6L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 4L, 4L,
4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 5L, 5L, 5L, 6L,
6L, 6L, 3L, 4L, 3L, 4L, 5L, 6L, 3L, 4L, 5L, 6L, 3L, 4L, 5L,
6L, 1L, 6L, 3L, 3L, 4L, 4L, 5L), .Label = c("AKT1", "ERBB2",
"ESR1", "PIK3CA", "TP53", "WT"), class = "factor"), LABO_ID = structure(c(45L,
8L, 13L, 11L, 11L, 26L, 7L, 15L, 23L, 26L, 35L, 39L, 7L,
19L, 26L, 32L, 33L, 35L, 39L, 15L, 19L, 35L, 1L, 37L, 34L,
43L, 47L, 3L, 10L, 18L, 20L, 28L, 31L, 36L, 42L, 9L, 10L,
14L, 18L, 20L, 28L, 31L, 36L, 44L, 45L, 8L, 10L, 18L, 28L,
42L, 2L, 7L, 39L, 7L, 39L, 3L, 4L, 42L, 5L, 42L, 6L, 21L,
1L, 10L, 22L, 28L, 46L, 9L, 10L, 14L, 28L, 46L, 10L, 28L,
48L, 25L, 23L, 32L, 33L, 40L, 43L, 24L, 3L, 18L, 24L, 28L,
31L, 36L, 42L, 18L, 27L, 28L, 31L, 36L, 45L, 18L, 24L, 27L,
28L, 42L, 16L, 16L, 18L, 18L, 18L, 29L, 23L, 39L, 39L, 40L,
1L, 12L, 47L, 3L, 18L, 20L, 28L, 31L, 36L, 38L, 42L, 5L,
18L, 20L, 27L, 28L, 31L, 36L, 38L, 41L, 45L, 8L, 18L, 27L,
28L, 42L, 48L, 6L, 17L, 30L, 31L, 31L, 18L, 18L, 18L, 29L,
39L, 39L, 40L, 43L, 31L, 31L, 48L, 30L, 13L, 34L, 18L, 36L,
18L, 36L, 18L), .Label = c("ER-11", "ER-19", "ER-21", "ER-22",
"ER-29", "ER-30", "ER-31", "ER-32", "ER-33", "ER-38", "ER-40",
"ER-43", "ER-49", "ER-8", "ER-AZ-04", "ER-AZ-05", "ER-AZ-06",
"ER-AZ-07", "ER-AZ-08", "ER-AZ-10", "ER-AZ-11", "ER-AZ-11=ER-47",
"ER-AZ-13", "ER-AZ-14", "ER-AZ-15", "ER-AZ-16", "ER-AZ-17",
"ER-AZ-18", "ER-AZ-20", "ER-AZ-20=ER-27", "ER-AZ-21", "ER-AZ-23",
"ER-AZ-23=ER-52", "ER-AZ-24", "ER-AZ-29", "ER-AZ-31", "ER-AZ-33",
"ER-AZ-35", "ER-AZ-37", "ER-AZ-38", "ER-AZ-39", "ER-AZ-40",
"ER-AZ-43", "ER-AZ-44", "ER-AZ-45", "ER-AZ-49", "ER-AZ-51",
"ER-AZ-53"), class = "factor"), Freq = c(1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L)), class = c("grouped_df", "tbl_df",
"tbl", "data.frame"), row.names = c(NA, -161L), groups = structure(list(
Metastasis_Location = c(1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L, 3L, 3L, 3L,
4L, 4L, 4L, 4L, 4L, 4L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 6L,
6L, 6L, 6L, 6L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 7L, 8L,
8L, 9L, 9L, 9L, 10L, 10L, 10L, 10L, 10L, 10L, 10L, 10L, 10L,
11L, 11L, 11L, 11L, 11L), T0_T2_THERAPY_COD = structure(c(2L,
2L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L,
2L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 1L, 1L, 2L, 2L, 2L, 2L, 1L,
1L, 1L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 2L, 2L, 2L, 1L, 1L, 1L,
1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 1L,
1L, 2L, 2L, 2L, 2L, 2L, 1L, 2L, 2L, 2L, 2L), .Label = c("A",
"F"), class = "factor"), T0_T2_PD_event = structure(c(2L,
2L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 1L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 2L, 2L,
2L, 2L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 1L, 2L, 2L,
2L, 1L, 2L, 2L, 2L, 2L, 1L, 1L, 2L, 2L, 2L), .Label = c("No Progression",
"Progression"), class = "factor"), Gene_mut = structure(c(4L,
5L, 1L, 3L, 4L, 1L, 2L, 3L, 4L, 5L, 6L, 3L, 6L, 3L, 4L, 5L,
6L, 2L, 3L, 4L, 3L, 4L, 5L, 6L, 3L, 6L, 3L, 4L, 5L, 6L, 3L,
4L, 5L, 6L, 1L, 3L, 4L, 5L, 3L, 4L, 3L, 4L, 5L, 6L, 3L, 4L,
5L, 6L, 6L, 3L, 4L, 5L, 6L, 3L, 4L, 3L, 4L, 5L, 6L, 3L, 4L,
5L, 6L, 3L, 4L, 5L, 6L, 1L, 6L, 3L, 4L, 5L), .Label = c("AKT1",
"ERBB2", "ESR1", "PIK3CA", "TP53", "WT"), class = "factor"),
.rows = structure(list(1L, 2L, 3L, 4L, 5L, 6L, 7L, 8:12,
13:19, 20:22, 23L, 24L, 25:27, 28:35, 36:45, 46:50, 51L,
52L, 53L, 54:55, 56:58, 59L, 60L, 61L, 62L, 63L, 64:67,
68:72, 73:75, 76L, 77L, 78:79, 80L, 81L, 82L, 83:89,
90:95, 96:100, 101L, 102L, 103L, 104L, 105L, 106L, 107:108,
109L, 110L, 111:112, 113L, 114:121, 122:131, 132:137,
138:140, 141L, 142L, 143L, 144L, 145L, 146L, 147L, 148L,
149L, 150L, 151L, 152L, 153L, 154L, 155L, 156L, 157:158,
159:160, 161L), ptype = integer(0), class = c("vctrs_list_of",
"vctrs_vctr", "list"))), class = c("tbl_df", "tbl", "data.frame"
), row.names = c(NA, -72L), .drop = TRUE))
CodePudding user response:
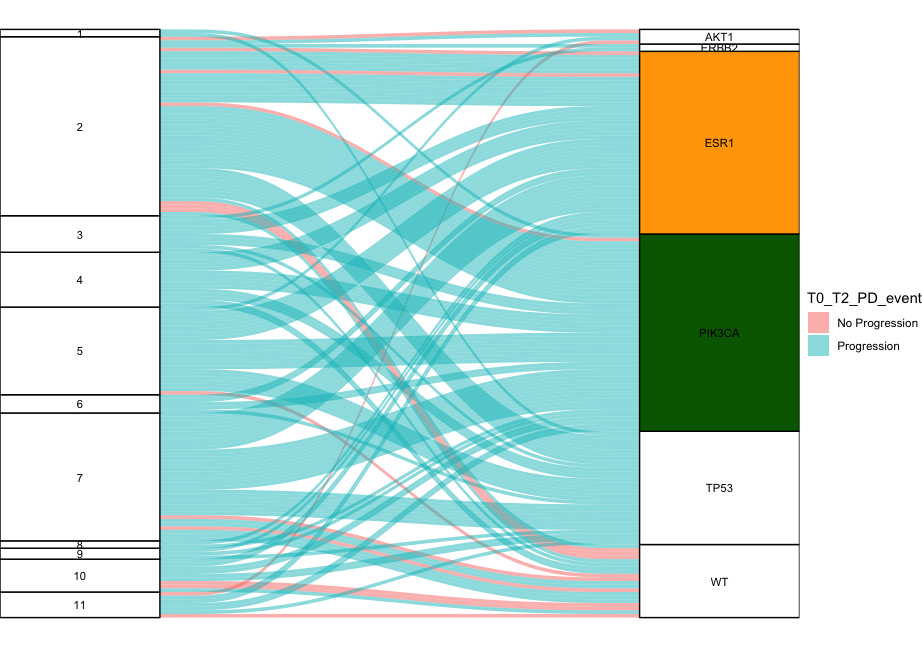
Here is a way you can get the colors - although someone will probably propose a more elegant way of writing it. You can change the colors in the first line of code if you don't like them.
library(ggalluvial)
library(ggplot2)
colorfill <- c("white", "white", "white", "white", "white", "white", "white", "white", "white", "white", "white", "white", "white", "darkgreen", "orange", "white", "white")
ggplot(data = Allu,
aes(axis1 = Metastasis_Location, axis2 = Gene_mut, y = Freq))
geom_alluvium(aes(fill = T0_T2_PD_event),
curve_type = "quintic")
geom_stratum(width = 1/4, fill = colorfill)
geom_text(stat = "stratum", size = 3,
aes(label = after_stat(stratum)))
scale_x_discrete(limits = c("Metastasis_Location", "Gene_mut"),
expand = c(0.05, .05))
theme_void()