I'm using a function to calculate the length of linestring per cell by ID and store in a list, convert each element of the list into a RasterLayer and turn that list into a RasterStack, average all layers and get a single raster.
#function
# build_length_raster <- function(one_df) {
intersect_list <- by(
one_df ,
one_df$sub_id,
function(subid_df) sf::st_intersection(grid2, subid_df) %>%
dplyr::mutate(length = as.numeric(sf::st_length(.))) %>%
sf::st_drop_geometry()
)
list_length_grid <- purrr::map(intersect_list, function(x)
x %>% dplyr::left_join(x=grid2, by="cell", copy=T) %>%
dplyr::mutate(length=length) %>%
dplyr::mutate_if(is.numeric,coalesce,0)
)
list_length_raster <- purrr::map(list_length_grid, function(x)
raster::rasterize(x, r, field="length", na.rm=F, background=0)
)
list_length_raster2 <- unlist(list_length_raster, recursive=F)
raster_stack <- raster::stack(list_length_raster2)
raster_mean <- raster::stackApply(
raster_stack,
indices = rep(1,nlayers(raster_stack)),
fun = "mean", na.rm = TRUE)
#}
The function presents a step where, in order for the resulting grid of st_intersection() to have the same number of cells as it had initially, I use left_join(by="cell" column).Then I use mutate() to replace the NA's with 0. When I run the function steps for one dataframe from the list, it works perfectly, but when I put it inside map() to do this in a list, I get this error, which seems to refer to the dplyr functions:
final_list <- purrr::map(mylist, build_length_raster)
> rlang::last_error()
<error/rlang_error>
Join columns must be present in data.
x Problem with `cell`.
Backtrace:
1. purrr::map(mylist, build_length_raster)
15. dplyr:::left_join.data.frame(., x = grid, by = "cell", copy = T)
16. dplyr:::join_mutate(...)
17. dplyr:::join_cols(...)
18. dplyr:::standardise_join_by(by, x_names = x_names, y_names = y_names)
19. dplyr:::check_join_vars(by$y, y_names)
Run `rlang::last_trace()` to see the full context.
Is there a way to solved this problem?
MYDATA example
library(tidyverse)
library(sf)
library(purrr)
library(raster)
#data example
id <- c("844", "844", "844", "844", "844","844", "844", "844", "844", "844",
"844", "844", "845", "845", "845", "845", "845","845", "845", "845",
"845","845", "845", "845")
sub_id <- c("2017_844_1", "2017_844_1", "2017_844_1", "2017_844_1", "2017_844_2",
"2017_844_2", "2017_844_2", "2017_844_2", "2017_844_3", "2017_844_3",
"2017_844_3", "2017_844_3", "2017_845_1", "2017_845_1", "2017_845_1",
"2017_845_1", "2017_845_2","2017_845_2", "2017_845_2", "2017_845_2",
"2017_845_3","2017_845_3", "2017_845_3", "2017_845_3")
lat <- c(-30.6456, -29.5648, -27.6667, -31.5587, -30.6934, -29.3147, -23.0538,
-26.5877, -26.6923, -23.40865, -23.1143, -23.28331, -31.6456, -24.5648,
-27.6867, -31.4587, -30.6784, -28.3447, -23.0466, -27.5877, -26.8524,
-23.8855, -24.1143, -23.5874)
long <- c(-50.4879, -49.8715, -51.8716, -50.4456, -50.9842, -51.9787, -41.2343,
-40.2859, -40.19599, -41.64302, -41.58042, -41.55057, -50.4576, -48.8715,
-51.4566, -51.4456, -50.4477, -50.9937, -41.4789, -41.3859, -40.2536,
-41.6502, -40.5442, -41.4057)
df <- tibble(id = as.factor(id), sub_id = as.factor(sub_id), lat, long)
#converting to sf
df.sf <- df %>%
sf::st_as_sf(coords = c("long", "lat"), crs = 4326)
#creating grid
xy <- sf::st_coordinates(df.sf)
grid = sf::st_make_grid(sf::st_bbox(df.sf),
cellsize = .1, square = FALSE) %>%
sf::st_as_sf()
#creating raster
r <- raster::raster(grid, res=0.1)
#return grid because raster function changes number of cells
grid2 <- rasterToPolygons(r, na.rm=F) %>%
st_as_sf() %>% mutate(cell=1:ncell(r))
#creating linestring to each sub_id
df.line <- df.sf %>%
dplyr::group_by(sub_id, id) %>%
dplyr::summarize() %>%
sf::st_cast("LINESTRING")
#creating ID list
mylist<- split(df.line, df.line$id)
#separating one dataframe of list to test function
one_df <- df.line[df.line$id=="844",]
one_df$id <- droplevels(one_df$id)
one_df$sub_id <- droplevels(one_df$sub_id)
CodePudding user response:
The specific error is caused because intersect_list has empty items in the list, which cannot be joined because they are empty, and hence have no columns to join by. If you modified the map function to only use non-empty items of intersect_list you would not get that error.
As you noted in the comments, removing the empty list entries with keep(intersect_list, ~ !is.null(.)) before mapping left_join onto the list items will fix the error.
However, I don't think this is the most elegant way to solve this problem. I might misunderstand what the goal is, but if it's to produce a raster from the total length of lines within each grid cell, I think a simpler approach without using purrr might work.
This is not the exact same as your product, but I'm keeping it simpler rn to illustrate an alternate approach. Here is a sum of the lengths in each cell as a stars object (similar to raster but plays better with the tidyverse and sf).
I'm starting off from your objects one_df and grid:
# Turn multiple lines into single MULTILINESTRING:
one_df %>%
st_union() ->
union_df
# Intersection of each grid cell with the MULTILINESTRING geometry:
grid %>%
st_intersection(union_df) ->
grid_lines
# Get lengths:
grid_lines %>%
mutate(length = st_length(x)) %>%
st_drop_geometry() ->
grid_lengths
# Join the calculated lengths back with the spatial grid,
# most of which will have NA for length
grid %>%
left_join(grid_lengths, by = "cell") ->
grid_with_lengths
# Rasterize the length field of the grid
grid_with_lengths %>%
dplyr::select(length) %>%
stars::st_rasterize() ->
length_stars
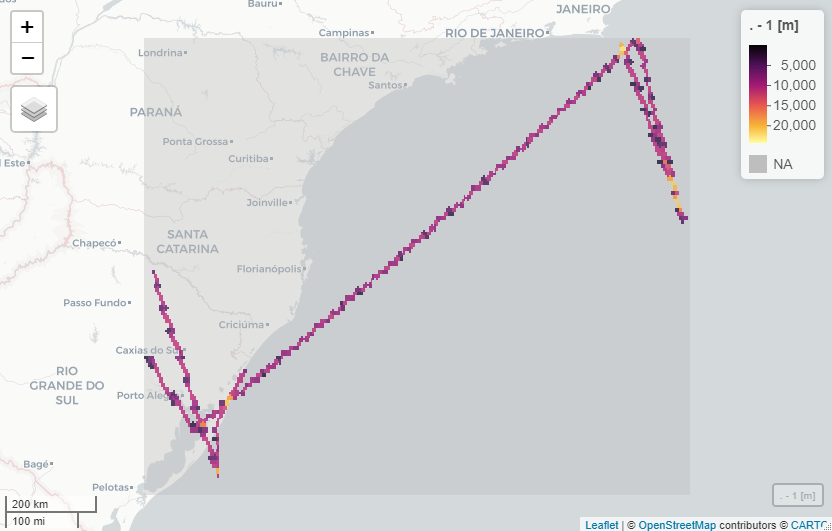
length_stars %>% mapview::mapview()