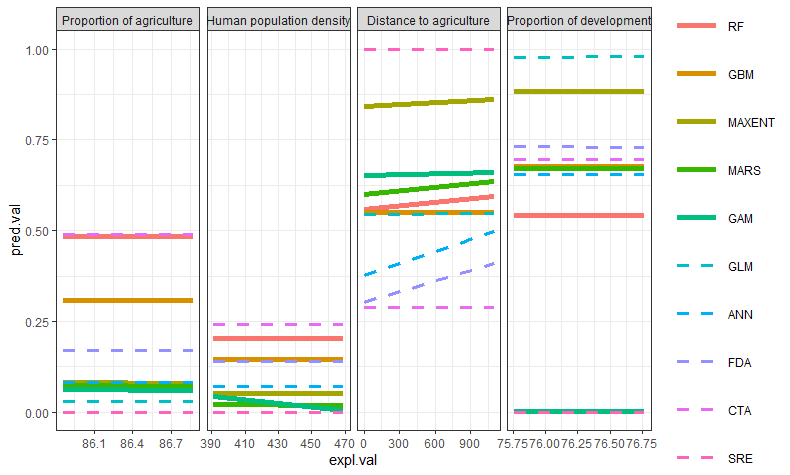
I have a plot with 5 lines that are dashed, 5 that are solid. This is correctly displaying in the plot. However, using the following code, the legend shows all line to be solid. I find this strange, because the legend is showing the correct variation in the line size, which is formatted in the same way. I tried replacing "Factorlevel" = "solid" by "Factorlevel" = 1, but this didn't work either.
Please see reproducible example below my code
ggplot(respplot_new, aes(x=expl.val,y=pred.val,color=model))
geom_line(aes(size=model,linetype=model))
scale_linetype_manual(values = c("RF" = "solid", "GBM" = "solid",
"MAXENT" = "solid", "MARS" = "solid", "GAM" = "solid",
"GLM"="dashed", "ANN"="dashed", "FDA"="dashed","CTA"="dashed","SRE"="dashed"))
scale_size_manual(values = c("RF" = 1.8, "GBM" = 1.8,
"MAXENT" = 1.8, "MARS" = 1.8, "GAM" = 1.8,
"GLM"=1.1, "ANN"=1.1, "FDA"=1.1,"CTA"=1.1,"SRE"=1.1))
facet_grid(~respplot_new$expl.name,scales="free") theme_bw()
Sorry for the length of the reproducible example, this is the minimum I was able to obtain with the dput function. Thank you for your consideration.
structure(list(expl.name = structure(c(3L, 3L, 3L, 3L, 3L, 3L,
3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L,
4L, 4L, 4L, 4L, 4L, 4L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L,
2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L), .Label = c("Proportion of agriculture",
"Human population density", "Distance to agriculture", "Proportion of development"
), class = "factor"), expl.val = c(0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 1098.78377525253, 1098.78377525253, 1098.78377525253, 1098.78377525253,
1098.78377525253, 1098.78377525253, 1098.78377525253, 1098.78377525253,
1098.78377525253, 1098.78377525253, 85.8585858585859, 85.8585858585859,
85.8585858585859, 85.8585858585859, 85.8585858585859, 85.8585858585859,
85.8585858585859, 85.8585858585859, 85.8585858585859, 85.8585858585859,
86.8686868686869, 86.8686868686869, 86.8686868686869, 86.8686868686869,
86.8686868686869, 86.8686868686869, 86.8686868686869, 86.8686868686869,
86.8686868686869, 86.8686868686869, 75.7575757575758, 75.7575757575758,
75.7575757575758, 75.7575757575758, 75.7575757575758, 75.7575757575758,
75.7575757575758, 75.7575757575758, 75.7575757575758, 75.7575757575758,
76.7676767676768, 76.7676767676768, 76.7676767676768, 76.7676767676768,
76.7676767676768, 76.7676767676768, 76.7676767676768, 76.7676767676768,
76.7676767676768, 76.7676767676768, 390.966472326806, 390.966472326806,
390.966472326806, 390.966472326806, 390.966472326806, 390.966472326806,
390.966472326806, 390.966472326806, 390.966472326806, 390.966472326806,
469.159766792168, 469.159766792168, 469.159766792168, 469.159766792168,
469.159766792168, 469.159766792168, 469.159766792168, 469.159766792168,
469.159766792168, 469.159766792168), model = structure(c(7L,
9L, 8L, 5L, 2L, 6L, 4L, 3L, 1L, 10L, 7L, 9L, 8L, 5L, 2L, 6L,
4L, 3L, 1L, 10L, 7L, 9L, 8L, 5L, 2L, 6L, 4L, 3L, 1L, 10L, 7L,
9L, 8L, 5L, 2L, 6L, 4L, 3L, 1L, 10L, 7L, 9L, 8L, 5L, 2L, 6L,
4L, 3L, 1L, 10L, 7L, 9L, 8L, 5L, 2L, 6L, 4L, 3L, 1L, 10L, 7L,
9L, 8L, 5L, 2L, 6L, 4L, 3L, 1L, 10L, 7L, 9L, 8L, 5L, 2L, 6L,
4L, 3L, 1L, 10L), .Label = c("RF", "GBM", "MAXENT", "MARS", "GAM",
"GLM", "ANN", "FDA", "CTA", "SRE"), class = "factor"), pred.val = c(0.377837081657681,
0.289682245014943, 0.303656725695349, 0.652763823576286, 0.551049518656044,
0.543664244044848, 0.600094628037518, 0.842622666666667, 0.557333333333333,
1, 0.497161495216137, 0.289682245014943, 0.411028449541226, 0.661018388330178,
0.550904016356143, 0.546817382114225, 0.634881521097495, 0.862494333333333,
0.594, 1, 0.0810608869754512, 0.489491284870227, 0.171208708485626,
0.0637801802410929, 0.307426323707738, 0.0308128470877357, 0.0743518990932153,
0.0817215, 0.484, 0, 0.0807936474564006, 0.489491284870227, 0.170148227471718,
0.0597404545728378, 0.307426323707738, 0.0295449629590053, 0.0716436251791972,
0.0804373, 0.483333333333333, 0, 0.654393286840172, 0.696277149376623,
0.73174123479955, 0.0033385823708866, 0.676035380732291, 0.978015696055508,
0.671377674947882, 0.883775666666667, 0.542, 0, 0.654393271023166,
0.696277149376623, 0.730485340340835, 0.00321040440788555, 0.676035380732291,
0.979003084409959, 0.670006581468528, 0.882102666666667, 0.542,
0, 0.0713256037593483, 0.242886821072384, 0.140090432179967,
0.042605020907364, 0.145155448037101, 2.42353541255428e-10, 0.0205399878748643,
0.0508441666666667, 0.204, 0, 0.0713256037593483, 0.242886821072384,
0.138548917887711, 0.00555445488463794, 0.145155448037101, 3.25156380242609e-12,
0.0187553675668009, 0.0508441666666667, 0.202666666666667, 0)), row.names = c(NA,
-80L), class = c("tbl_df", "tbl", "data.frame"))
CodePudding user response:
You may use theme(legend.key.size = unit(0.5, "in")) to increase size of legend, that have enough space to dashed line looks like dashed.
ggplot(respplot_new, aes(x=expl.val,y=pred.val,color=model))
geom_line(aes(size=model,linetype=model))
scale_linetype_manual(values = c("RF" = "solid", "GBM" = "solid",
"MAXENT" = "solid", "MARS" = "solid", "GAM" = "solid",
"GLM"="dashed", "ANN"="dashed", "FDA"="dashed","CTA"="dashed","SRE"="dashed"))
scale_size_manual(values = c("RF" = 1.8, "GBM" = 1.8,
"MAXENT" = 1.8, "MARS" = 1.8, "GAM" = 1.8,
"GLM"=1.1, "ANN"=1.1, "FDA"=1.1,"CTA"=1.1,"SRE"=1.1))
facet_grid(~respplot_new$expl.name,scales="free") theme_bw() theme(legend.key.size = unit(0.5, "in"))