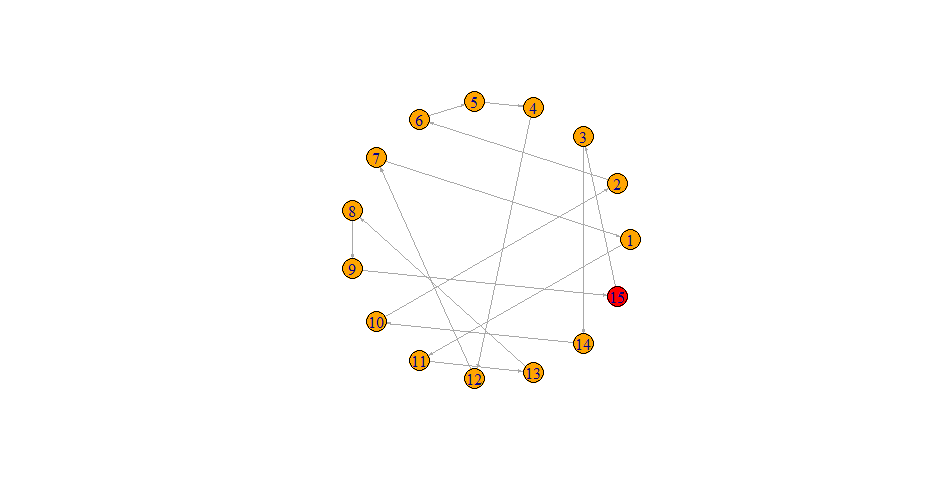
I have the following network graph:
library(tidyverse)
library(igraph)
set.seed(123)
n=15
data = tibble(d = paste(1:n))
relations = tibble(
from = sample(data$d),
to = lead(from, default=from[1]),
)
graph = graph_from_data_frame(relations, directed=T, vertices = data)
V(graph)$color <- ifelse(data$d == relations$from[1], "red", "orange")
plot(graph, layout=layout.circle, edge.arrow.size = 0.2)
I learned how to remove all the "edges" in this graph:
g <- graph-E(graph)
plot(g)
Now, I am trying to do this same thing, but using a "loop" instead:
for (i in 1:15)
for (j in 1:15) {
graph <- graph - edge(paste0(i, "|", j))
}
But I think the problem is that the above code is trying to delete "edges" that exist, and this code does not have the ability to "skip" (override) an instance when it is commanded to delete a non-existing edge:
Error in delete_edges(e1, unlist(e2, recursive = FALSE)) :
At iterators.c:1828 : Cannot create iterator, invalid edge id, Invalid vertex id
Is there a way to instruct this "loop" to "skip" every instances where two edges do not have a connection and to continue until the loop is done?
Thank you!
CodePudding user response:
I don't know why you want to run a for loop, but please find below a possible solution using the as_edgelist() function from the igraph library.
Reprex
- Your data
library(igraph)
library(dplyr)
set.seed(123)
n=15
data = tibble(d = paste(1:n))
relations = tibble(
from = sample(data$d),
to = lead(from, default=from[1]),
)
graph = graph_from_data_frame(relations, directed=T, vertices = data)
V(graph)$color <- ifelse(data$d == relations$from[1], "red", "orange")
plot(graph, layout=layout.circle, edge.arrow.size = 0.2)
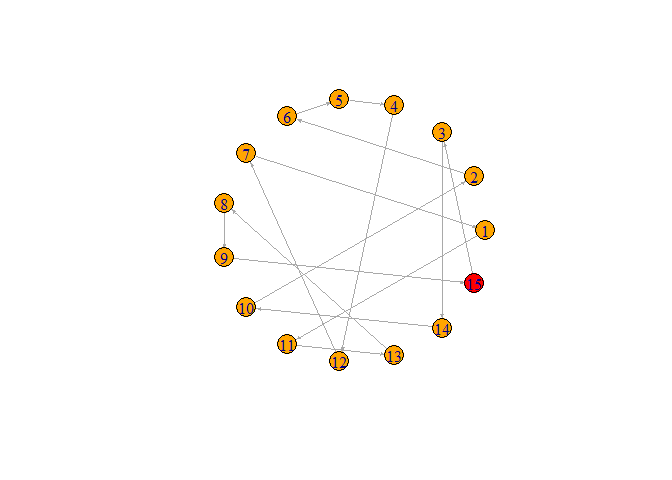
- Suggested code
edgelist <- as_edgelist(graph, names = TRUE)
for (i in 1:15) {
graph <- graph - edge(paste0(edgelist[i,1], "|", edgelist[i,2]))
}
plot(graph)
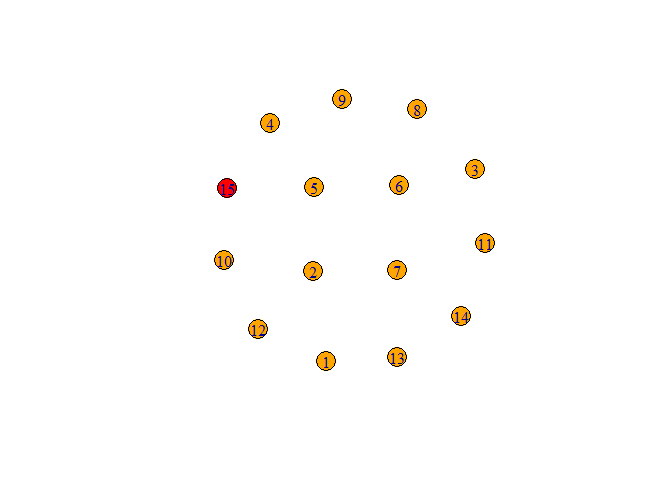

Created on 2022-02-24 by the reprex package (v2.0.1)