I am using R studio. What happens is that I have a dataset in which I have 1000k data. I have all columns called FINAL_CLASSIFICATION and AGE. In the FINAL_RANKING column there is data ranging from 1 to 7. In this column we say that those with 1, 2 or 3, are infected with SARS_COVID, while in those with 4, 5,6 and 7 are those who are healthy. I need to make a histogram of the ages of those who are infected and for this I understand that I must make a group to see the ages that coincide with 1, 2 and 3 of the column CLASIFICACION_FINAL and those ages will be of the infected people and, from there I need to make the histogram but I do not find the way to create the group or to obtain this.
Could you help me?
I have the following code
#1)
# import the data into R
# RECOMMENDATION: use read_csv
covid_dataset <- read_csv("Desktop/Course in R/Examples/covid_dataset.csv")
View(covid_dataset)
#------------------------------------------------------------------------------------------
#2) Extract a random sample of 100k records and assign it into a new variable. From now on work with this dataset
# HINT: use dplyr's sample_n function
sample <- sample_n(covid_dataset, 100000)
# With the function sample_n what we get is a syntax sample_n(x,n) where we have that
#x will be our dataset from where we want to extract the sample and n is the sample size
#that we want
nrow(sample)
#with this function we can corroborate that we have extracted a 100K sample.
#------------------------------------------------------------------------------------------
#3)Make a statistical summary of the dataset and also show the data types by column.
summary(sample)
#The summary function is the one that gives us the summary statistics.
map(sample, class)
#The map() function gives us the data type by columns and we can see that there are
#more numeric data type.
#-------------------------------------------------------------------------------------------
#4)Filter the rows that are positive for SARS-COVID and calculate the number of records.
## Positive cases are those that in the FINAL_CLASSIFICATION column have 1, 2 or 3.
## To filter the rows, we will make use of the PIPE operator and the select function of dplyr.
#This will help us to select the column and to be able to filter the rows where
#the FINAL_CLASSIFICATION column is 1, 2 or 3, i.e. SARS-COVID positive results.
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION == 1)
# Here I filter the rows for which the column FINAL_CLASSIFICATION has a 1
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION == 2)
# Here I filter the rows for which the column FINAL_CLASSIFICATION has a 2
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION == 3)
# Here I filter the rows for which the column FINAL_CLASSIFICATION has a 3
# I do them separately to have a better view of the records.
#Now if we want to get them all together we simply do the following
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION <= 3)
#This gives us the rows less than or equal to 3, which is the same as giving us the rows in which the
#Rows where the FINAL_RANKING column has 1, 2 or 3.
#Now, if we want the number of records, doing it separately, we simply add
#another PIPE operator in which we will add the nrow() function to give me the number of #rows for each record.
#rows for each record.
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION == 1) %>% nrow()
#gives us a result of 1471
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION == 2) %>% nrow()
#gives us a result of 46
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION == 3) %>% nrow()
#Gives us a result of 37703
#If we add the 3 results, we have that the total number of records is
1471 46 37703
#Which gives us 39220
#But it can be simplified by doing it in a straightforward way as follows
sample %>% select(FINAL_CLASSIFICATION) %>% filter(FINAL_CLASSIFICATION <= 3) %>% nrow()
#And we notice that we get the same result as the previous code.
#In conclusion, we have a total of 39220 positive SARS-COVID cases.
#---------------------------------------------------------------------------------------------
#5)Count the number of null records per column (HINT: Use sapply or map, and is.na)
apply(sample, MARGIN = 2, function(x)sum(is.na(x))))
#This shows us the number of NA's per column. We notice that the only column
#that has NA's is the DATE_DEF with a total of 95044, this tells us that out of the
#100K data, only approximately 5k data are known for DATE_DEF.
#------------------------------------------------------------------------------------------
#6)
##a)Calculate the mean age of covid infectees.
##b)Make a histogram of the ages of the infected persons.
##c)Make a density plot of the ages of the infected persons
sample %>% group_by(FINAL_CLASSIFICATION
group_by(FINAL_CLASSIFICATION <= 3 ) %>% %>%
summarise(average = mean(AGE))
#Then the total average number of infected is 43.9
#Now we make a histogram of the ages of the infected persons
sample %>% group_by(FINAL_CLASSIFICATION <=3, AGE) %>% summarise(count = n())
It is in the last part where I have doubts. I want to find the average of the ages of the infected people, I used the code that I put there using group_by but I don't know if that is correct. And my doubts are already with the other two questions in #6, where I want to know about the histograms and how to plot them.
CodePudding user response:
What I gathered is that you wish to 1. create a variable 'FINAL_CLASSIFICATION' based on values of 'FINAL_RANKING,' 2. summarize the average age of groups in FINAL_CLASSIFICATION, and 3. create a histogram of the positive cases in FINAL_CLASSIFICATION
I created a random sample of 100 cases with random assumptions of AGE and FINAL_RANKING
library(dplyr)
library(ggplot2)
sample <- tibble(FINAL_RANKING = sample(1:7, 100, replace = T), AGE = sample(10:100, 100, replace = T) )
sample <- sample %>%
mutate(
FINAL_CLASSIFICATION = case_when(
FINAL_RANKING %in% 1:3 ~ "SARS_COVID_POSITIVE",
FINAL_RANKING %in% 4:7 ~ "SARS_COVID_NEGATIVE")
)
sample %>%
group_by(FINAL_CLASSIFICATION) %>%
summarize(average_age = mean(AGE))
sample %>%
filter(FINAL_CLASSIFICATION == "SARS_COVID_POSITIVE") %>%
ggplot(., aes(x = AGE))
geom_histogram()
Gives summary output:
# A tibble: 2 x 2
FINAL_CLASSIFICATION average_age
<chr> <dbl>
1 SARS_COVID_NEGATIVE 51.8
2 SARS_COVID_POSITIVE 58.6
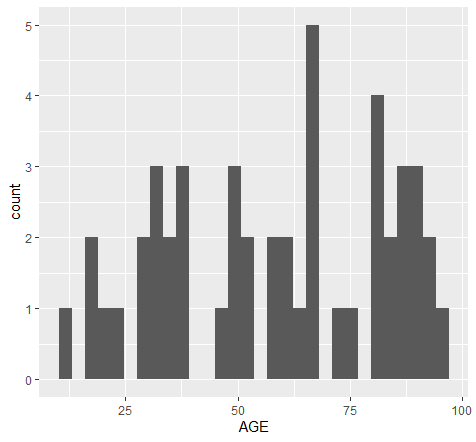
and plot:
As noted in output, you should adjust bins