I created a heatmap and a pca plot and tried to merge them as single figure. But the are not displayed as single figure.
library(factoextra)
library(FactoMineR)
library(pheatmap)
library(RColorBrewer)
library(ggpubr)
# make test matrix
test = matrix(rnorm(200), 20, 10)
test[1:10, seq(1, 10, 2)] = test[1:10, seq(1, 10, 2)] 3
test[11:20, seq(2, 10, 2)] = test[11:20, seq(2, 10, 2)] 2
test[15:20, seq(2, 10, 2)] = test[15:20, seq(2, 10, 2)] 4
colnames(test) = paste("Test", 1:10, sep = "")
rownames(test) = paste("Gene", 1:20, sep = "")
# define the annotation
annotation_row = data.frame(GeneClass = factor(rep(c("Path1", "Path2", "Path3"), c(10, 4, 6))),
AdditionalAnnotation = c(rep("random1", 10), rep("random2", 10)))
rownames(annotation_row) = paste("Gene", 1:20, sep = "")
a=pheatmap(test, annotation_row = annotation_row)
# creating pca
# Compute PCA with ncp = 3
res.pca <- PCA(test, ncp = 3, graph = FALSE)
# Compute hierarchical clustering on principal components
res.hcpc <- HCPC(res.pca, graph = FALSE)
# Principal components tree
b=plot(res.hcpc, choice = "3D.map")
#arranging in a single plot
ggarrange(a$gtable, b, labels = c("A", "B"))
CodePudding user response:
plot (or plot.HCPC) returns NULL and therefore b is NULL.
And from ?ggarrange, it expects a list of plots to be arranged into the grid. The plots can be either ggplot2 plot objects or arbitrary gtables.
So one option could be using as.ggplot() function from {ggplotify} package to convert that base plot to ggplot object and then pass it to ggarrange.
b <- ggplotify::as.ggplot(~plot(res.hcpc, choice = "3D.map"))
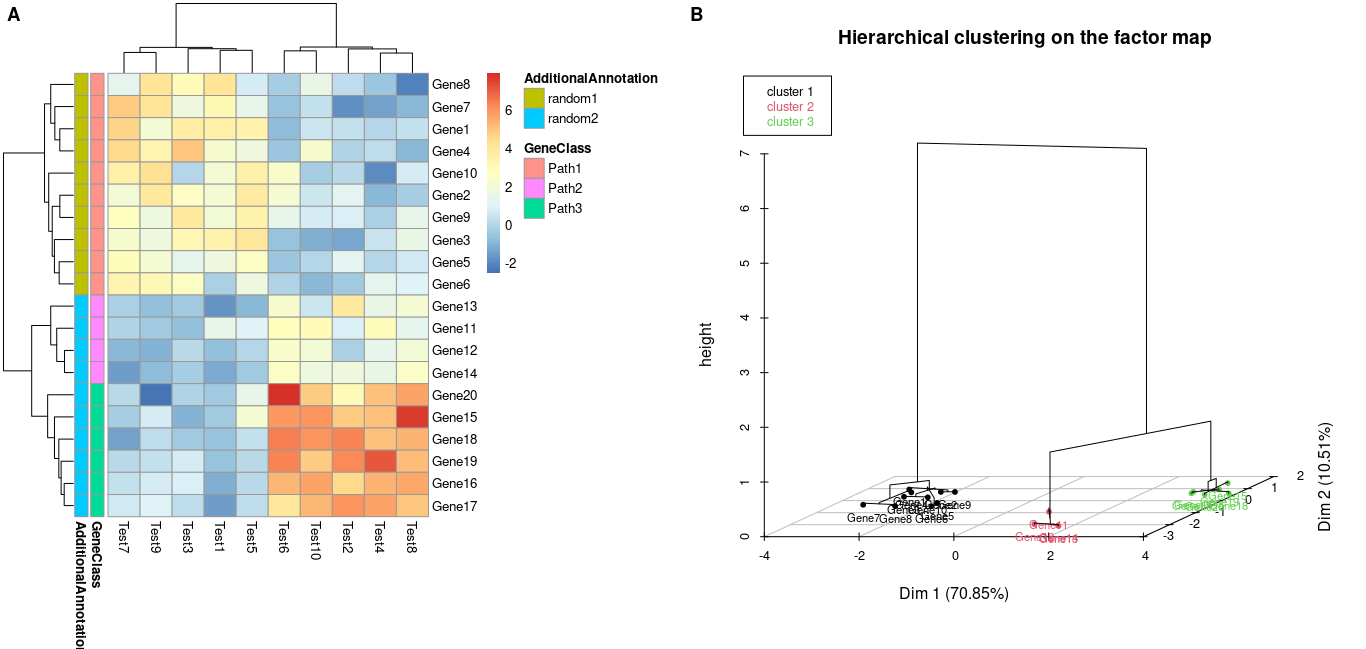
#arranging in a single plot
ggarrange(a$gtable, b, labels = c("A", "B"))