I would like to plot the proportion of levels of a group alongside a continuous variable. Since the x-axis is continuous, it is not really possible to compute proportions at each point (since there is an infinite number of them). So, usually, one cuts the continuous variable into bins, and plot them. Another solution is to use the density, but I want the proportions (so, the percentage) in the y-axis and I'm pretty sure density is not about proportions.
As an example, let's use iris and try to plot the share of each species among Sepal.Length. One can create bins using Hmisc::cut2 and then count the proportions for each group:
library(tidyverse)
library(Hmisc)
dat <- iris %>%
mutate(Sepal.Length = Sepal.Length rnorm(n()),
cut = cut2(Sepal.Length, g = 30, levels.mean = T)) %>%
group_by(cut) %>%
summarise(set = sum(Species == "setosa") / n(),
vir = sum(Species == "virginica") / n(),
ver = sum(Species == "versicolor") / n()) %>%
pivot_longer(-cut)
# A tibble: 90 x 3
cut name value
<fct> <chr> <dbl>
1 3.0126 set 0.6
2 3.0126 vir 0
3 3.0126 ver 0.4
4 3.7616 set 0.8
5 3.7616 vir 0
6 3.7616 ver 0.2
7 3.9898 set 0.8
8 3.9898 vir 0
9 3.9898 ver 0.2
10 4.1577 set 0.2
# ... with 80 more rows
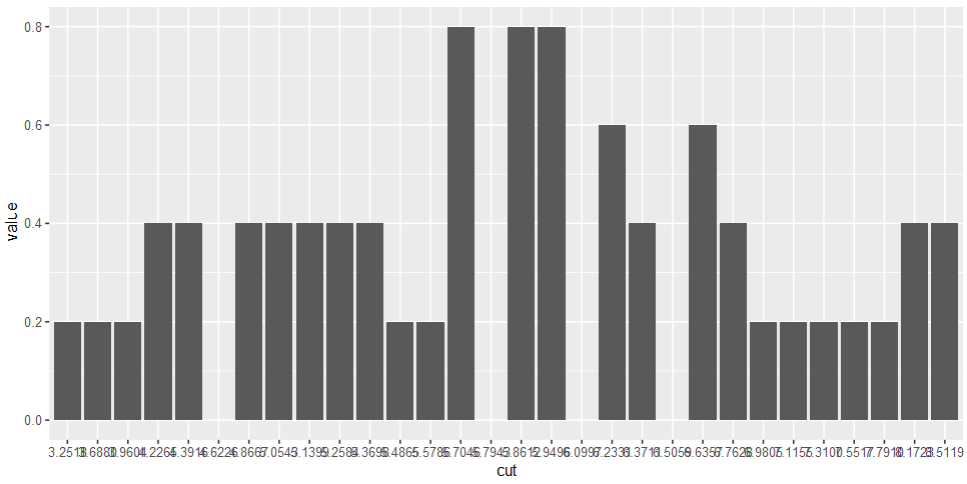
And the plot looks like this, e.g. for name == "ver"
dat %>%
filter(name == "ver") %>%
ggplot(aes(x = cut, y = value))
geom_col()
Now, is there any way to make this easier, and more esthetic?
Especially, making the x-axis a continuous back again so that one could e.g. create a geom_line between every columns of the plot (maybe making rolling means?). Or is it a bad practice and that's why I can't see any documentation about this?
CodePudding user response:
Setting the variable cut to numeric did the job, but there may still be better options.
dat %>%
filter(name == "ver") %>%
ggplot(aes(x = as.numeric(as.character(cut)), y = value))
geom_col()
Or with a line:
dat %>%
filter(name == "ver") %>%
ggplot(aes(x = as.numeric(as.character(cut)), y = value))
geom_line()