I am new to R and have the following example code that I wish to apply for every column in my data.
data(economics, package="ggplot2")
economics$index <- 1:nrow(economics)
loessMod10 <- loess(uempmed ~ index, data=economics, span=0.10)
smoothed10 <- predict(loessMod10)
plot(economics$uempmed, x=economics$date, type="l", main="Loess Smoothing and Prediction", xlab="Date", ylab="Unemployment (Median)")
lines(smoothed10, x=economics$date, col="red")
Could someone please suggest how this would be possible?
CodePudding user response:
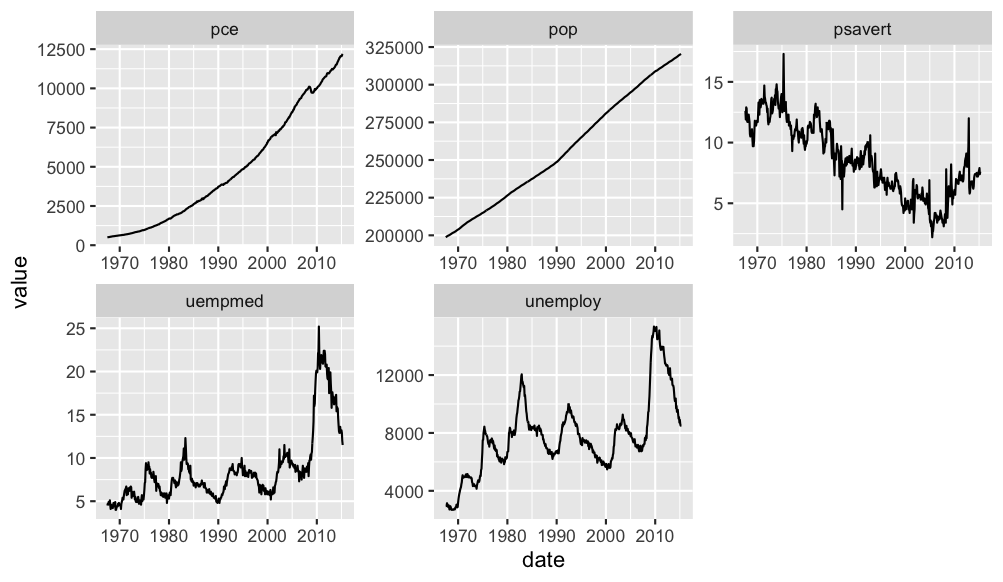
You can make your data from wide to long by the date and use facet_wrap. Maybe you want something like this:
library(ggplot2)
library(reshape2)
library(dplyr)
economics %>%
melt(., "date") %>%
ggplot(., aes(date, value))
geom_line()
facet_wrap(~variable, scales = "free")
Output:
Comment: All plots in one graph
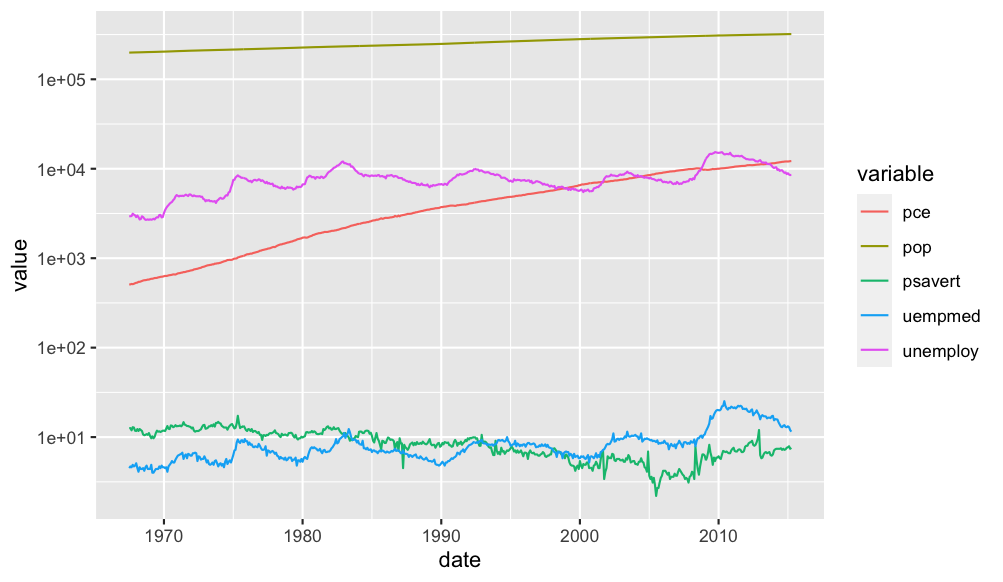
If you mean all plots in one graph, you can give the variables a color like this:
economics %>%
melt(., "date") %>%
ggplot(., aes(date, value, color = variable))
geom_line()
scale_y_log10()
Output:
CodePudding user response:
It's possible to perform loess smoothing within ggplot.
library(data.table)
library(ggplot2)
df <- economics
##
#
gg.melt <- setDT(df) |> melt(id='date', variable.name = 'KPI')
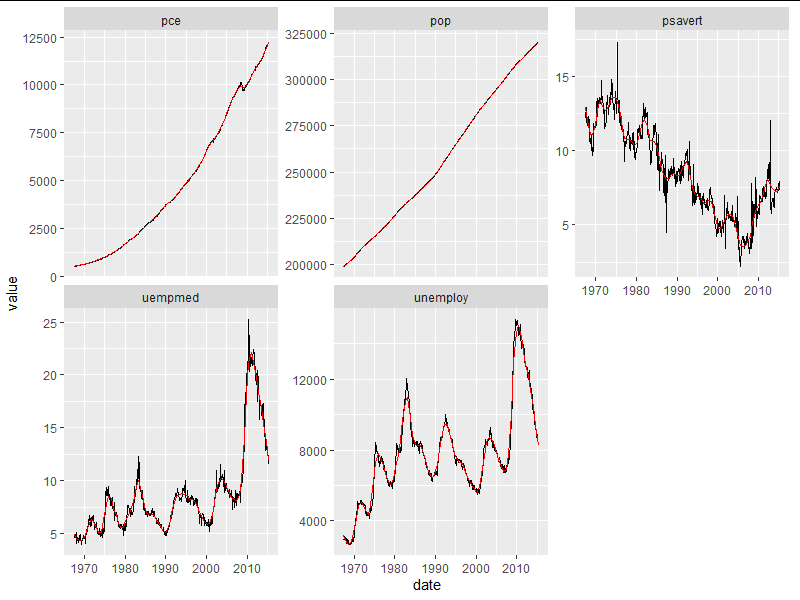
ggplot(gg.melt, aes(x=date, y=value))
geom_line()
stat_smooth(method=loess, color='red', size=0.5, se=FALSE, method.args = list(span=0.1))
facet_wrap(~KPI, scales = 'free_y')
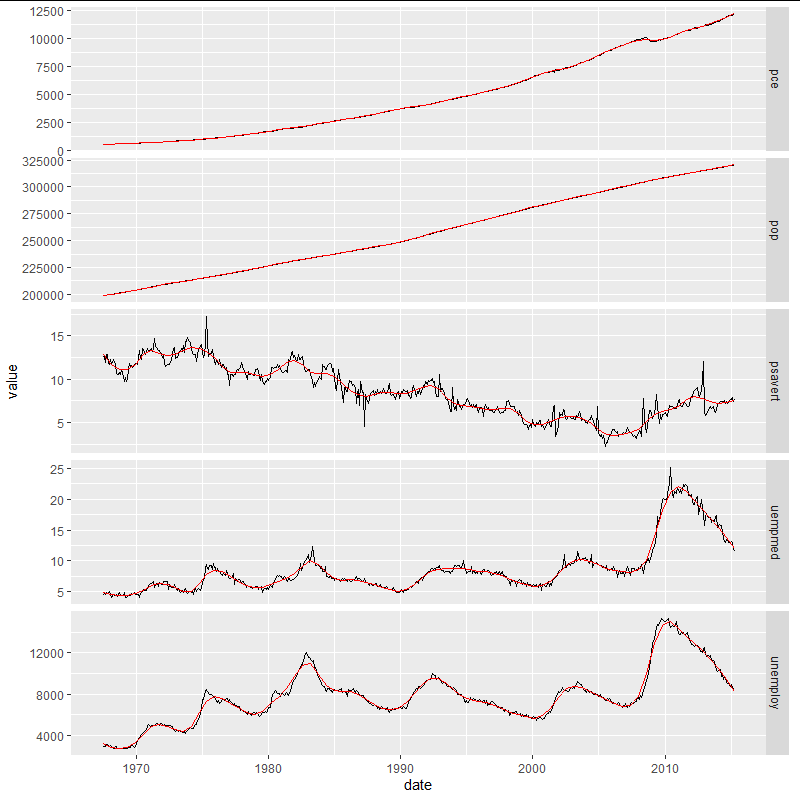
Regarding combining everything on one plot I'm not seeing how you would do that as the y-scales are so different. If the point is to see how the peaks line up, etc. you could do this:
ggplot(gg.melt, aes(x=date, y=value))
geom_line()
stat_smooth(method=loess, color='red', size=0.5, se=FALSE, method.args = list(span=0.1))
facet_grid(KPI~., scales = 'free_y')
There is also the dygraphs package which allows creation of dynamic graphics that can be saved to html:
gg.melt[, scaled:=scale(value, center = FALSE, scale=diff(range(value))), by=.(KPI)]
gg.melt[, pred:=predict(loess(scaled~as.integer(date), .SD, span=0.1)), by=.(KPI)]
gg.dt <- dcast(gg.melt, date~KPI, value.var = list('scaled', 'pred'))
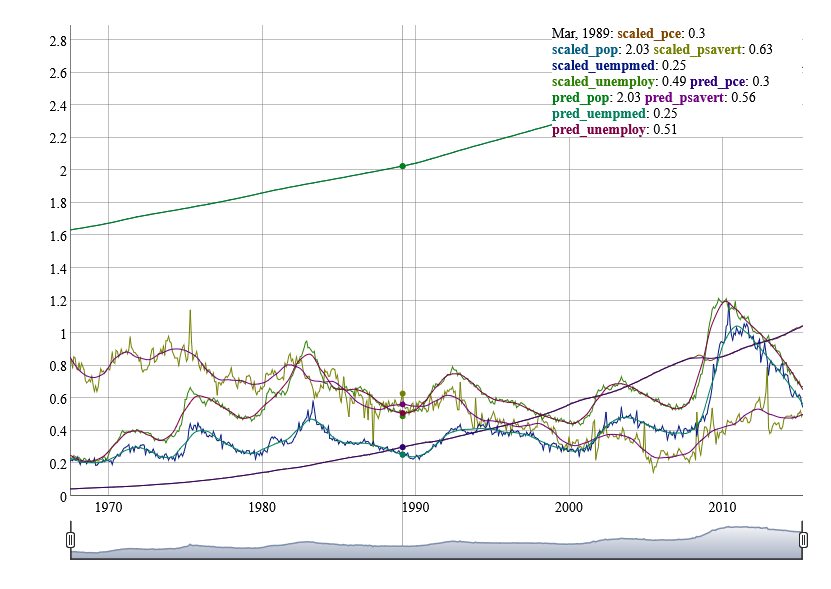
library(dygraphs)
dygraph(gg.dt) |>
dyCrosshair(direction = 'vertical') |>
dyRangeSelector()
It's possible to create a dygraph(...) version of the second plot, where the different KPI are in different facets, but you have to use RMarkdown for that.