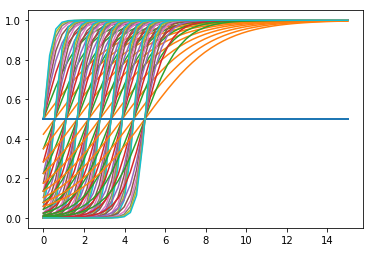
There are a few posts about this and normally the answer is to have a good initial guess and bounds. I've played around with it for a while and cannot find a configuration that produces any sort of curve.
import numpy as np
array1 = np.array(column1).astype(float)
array2 = np.array(column2).astype(float)
print(array1)
print(array2)
Output: [18.7327 9.3784 6.6293 20.8361 11.2603 19.3706 5.4302 10.1293 13.7516 8.0567 16.8688 4.969 3.94 19.4793 11.7527 13.2811 13.338 0.5944 7.4406 11.2338 6.2283 3.4818 10.1056 16.2689 22.442 18.7345 5.2605 5.6405 12.7186 18.2497 5.4315 14.2651 16.7544 12.9192 13.5955 10.9256 5.7798 8.4485 8.5229 11.879 6.5271 10.3376 7.781 31.4558 8.0236 2.3527 10.8926 16.1995 11.1924 25.8071 13.9692 20.7791 10.3045 12.2833 7.4066 15.9807 11.4462 15.1504 5.9021 19.1184]
[83.85 52.45 41.2 92.59 62.65 86.77 30.63 53.78 73.34 48.55 82.53 28.3 23.87 90.99 62.95 68.82 71.06 20.74 45.25 60.65 39.07 21.93 53.35 79.61 93.27 85.88 28.95 32.73 65.89 83.51 30.74 75.22 79.8 67.43 71.12 58.41 35.83 49.61 50.72 63.49 40.67 55.75 46.49 96.22 47.62 21.8 56.23 76.97 59.07 94.67 74.9 92.52 55.61 63.51 41.34 76.8 62.81 75.99 36.34 85.96]
import pylab
from scipy.optimize import curve_fit
def sigmoid(x, a, b):
y = 1 / (1 np.exp(-b*(x-a)))
return y
popt, pcov = curve_fit(sigmoid, array1, array2, p0 = [5,20], method='dogbox', bounds=([0, 20],[40, 100]))
print(popt)
x = np.linspace(0, 35, 50)
y = sigmoid(x, *popt)
pylab.plot(array1, array2, 'o', label='data')
pylab.plot(x,y, label='fit')
pylab.ylim(0, 100)
pylab.legend(loc='best')
pylab.show()
Therefore, you either have to modify your model to accept a scaling parameter, or scale down your data to a range your model can fit. Here's the two solutions:
Preamble
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
import pandas as pd
array1 = np.array([18.7327,9.3784,6.6293,20.8361,11.2603,19.3706,5.4302,10.1293,13.7516,8.0567,16.8688,4.969,3.94,19.4793,11.7527,13.2811,13.338,0.5944,7.4406,11.2338,6.2283,3.4818,10.1056,16.2689,22.442,18.7345,5.2605,5.6405,12.7186,18.2497,5.4315,14.2651,16.7544,12.9192,13.5955,10.9256,5.7798,8.4485,8.5229,11.879,6.5271,10.3376,7.781,31.4558,8.0236,2.3527,10.8926,16.1995,11.1924,25.8071,13.9692,20.7791,10.3045,12.2833,7.4066,15.9807,11.4462,15.1504,5.9021,19.1184])
array2 = np.array([83.85,52.45,41.2,92.59,62.65,86.77,30.63,53.78,73.34,48.55,82.53,28.3,23.87,90.99,62.95,68.82,71.06,20.74,45.25,60.65,39.07,21.93,53.35,79.61,93.27,85.88,28.95,32.73,65.89,83.51,30.74,75.22,79.8,67.43,71.12,58.41,35.83,49.61,50.72,63.49,40.67,55.75,46.49,96.22,47.62,21.8,56.23,76.97,59.07,94.67,74.9,92.52,55.61,63.51,41.34,76.8,62.81,75.99,36.34,85.96])
df = pd.DataFrame({'x':array1, 'y':array2})
df = df.sort_values('x')
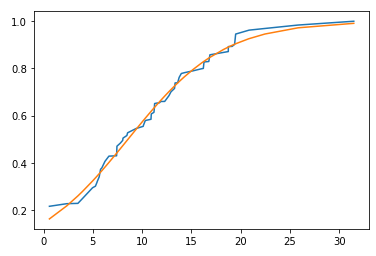
Scaling data to match parameter
def sigmoid(x, a, b):
y = 1 / (1 np.exp(-b*(x-a)))
return y
popt, pcov = curve_fit(sigmoid, df['x'], df['y'] / df['y'].max(), p0 = [5,20], method='dogbox', bounds=([0, 0],[40, 100]))
plt.plot(df['x'], df['y'] / df['y'].max(), label='data')
plt.plot(df['x'], sigmoid(df['x'], *popt))
popt is [8.56754823 0.20609918]
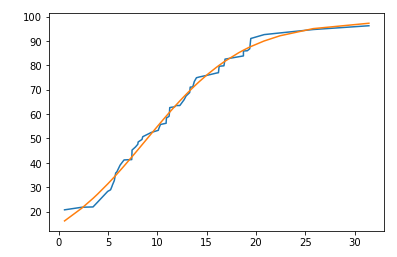
Adding new parameter to function
def sigmoid2(x, a, b, scale):
y = scale / (1 np.exp(-b*(x-a)))
return y
popt, pcov = curve_fit(sigmoid2, df['x'], df['y'], p0 = [5,20, 100], method='dogbox', bounds=([0, 0, 0],[40, 100, 1E5]))
plt.plot(df['x'], df['y'], label='data')
plt.plot(df['x'], sigmoid2(df['x'], *popt))
popt is array([ 8.81708442, 0.19749557, 98.357044 ])