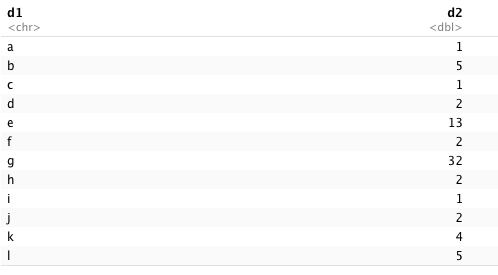
I have a data frame like below
d1<-c('a','b','c','d','e','f','g','h','i','j','k','l')
d2<-c(1,5,1,2,13,2,32,2,1,2,4,5)
df1<-data.frame(d1,d2)
Which looks like the data table in this picture
My goal is to filter the rows based on which value of d2 in every 3 rows is biggest. So it would look like this:
 Thank you!
Thank you!
CodePudding user response:
We may use rollmax from zoo to filter the rows
library(dplyr)
library(zoo)
df1 %>%
filter(d2 == na.locf0(rollmax(d2, k = 3, fill = NA)))
d1 d2
1 b 5
2 e 13
3 g 32
4 l 5
CodePudding user response:
You can create a grouping variable that puts observations into groups of 3. I have first created a sequence from 1 to the total number of rows, incremented by 3. And then repeated each number of this sequence 3 times and subset the result to get a vector the same length of the data, incase the number of observations is not perfectly divisible by 3. Then simply filter rows based by the largest number of each group in d2 column.
library(dplyr)
df1 %>%
mutate(group = rep(seq(1, n(), by = 3), each = 3)[1:n()]) %>%
group_by(group) %>%
filter(d2 == max(d2))
# A tibble: 4 x 3
# Groups: group [4]
# d1 d2 group
# <chr> <dbl> <dbl>
# 1 b 5 1
# 2 e 13 4
# 3 g 32 7
# 4 l 5 10
CodePudding user response:
Yet another solution:
library(tidyverse)
d1<-c('a','b','c','d','e','f','g','h','i','j','k','l')
d2<-c(1,5,1,2,13,2,32,2,1,2,4,5)
df1<-data.frame(d1,d2)
df1 %>%
mutate(id = rep(1:(n()/3), each=3)) %>%
group_by(id) %>%
slice_max(d2) %>%
ungroup %>% select(-id)
#> # A tibble: 4 × 2
#> d1 d2
#> <chr> <dbl>
#> 1 b 5
#> 2 e 13
#> 3 g 32
#> 4 l 5