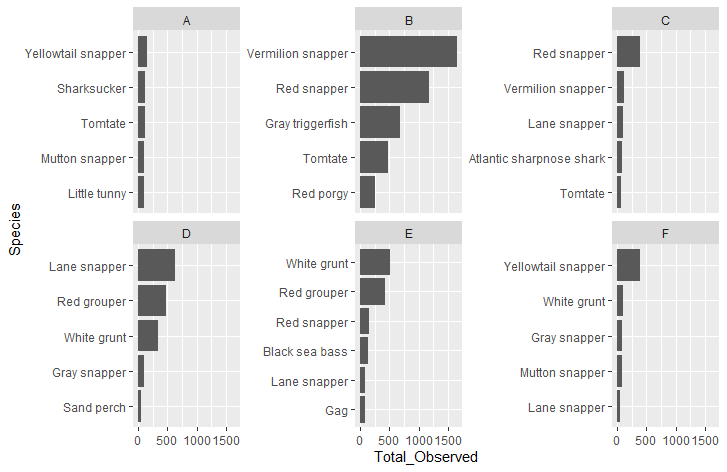
I am trying to display data by Species that has different values depending on group Letter. The best way I have found to display my data is by putting my categorical data on the y-axis and displaying the Total_Observed on the x-axis. Lemon allows me to have different y-axis labels. Unfortunately, the graph sorts by my y-axis labels instead of using my data as is, which is sorted by most abundant species to least abundant. Any suggestions?
Using libraries: dplyr, ggplot2, lemon
My data:
|Letter |Species | Total_Observed|
|:------|:------------------------|--------------:|
|A |Yellowtail snapper | 155|
|A |Sharksucker | 119|
|A |Tomtate | 116|
|A |Mutton snapper | 104|
|A |Little tunny | 96|
|B |Vermilion snapper | 1655|
|B |Red snapper | 1168|
|B |Gray triggerfish | 689|
|B |Tomtate | 477|
|B |Red porgy | 253|
|C |Red snapper | 391|
|C |Vermilion snapper | 114|
|C |Lane snapper | 95|
|C |Atlantic sharpnose shark | 86|
|C |Tomtate | 73|
|D |Lane snapper | 627|
|D |Red grouper | 476|
|D |White grunt | 335|
|D |Gray snapper | 102|
|D |Sand perch | 50|
|E |White grunt | 515|
|E |Red grouper | 426|
|E |Red snapper | 150|
|E |Black sea bass | 142|
|E |Lane snapper | 88|
|E |Gag | 88|
|F |Yellowtail snapper | 385|
|F |White grunt | 105|
|F |Gray snapper | 88|
|F |Mutton snapper | 82|
|F |Lane snapper | 59|
Then I run the code for my ggplot/lemon
ggplot(test,aes(y=Species,x=Total_Observed)) geom_histogram(stat='identity') facet_wrap(.~test$Letter,scales='free_y')
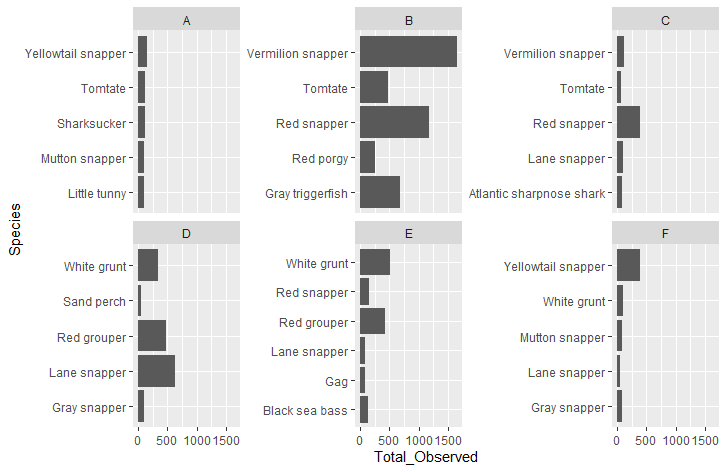
And my graphs print like this:
CodePudding user response: