I have this data with one column in character and one column in value.
data = structure(list(Station = c("1A", "1B", "2A", "2B", "3A", "3B",
"4A", "4B", "5A", "5B", "6A", "6B", "7A", "7B"), Particles.kg = c(370L,
420L, 250L, 320L, 130L, 210L, 290L, 390L, 230L, 340L, 60L, 90L,
130L, 170L)), class = "data.frame", row.names = c(NA, -14L))
now i convert the character in to factor by
data$Station = as.factor(data$Station)
then i start Hierarchical Cluster analysis
rownames(data) = c(data$Station)
data = data[,-1]
require(stats)
res.dist = dist(x=data, method = "euclidean")
hcl = hclust(d=res.dist, method = "complete")
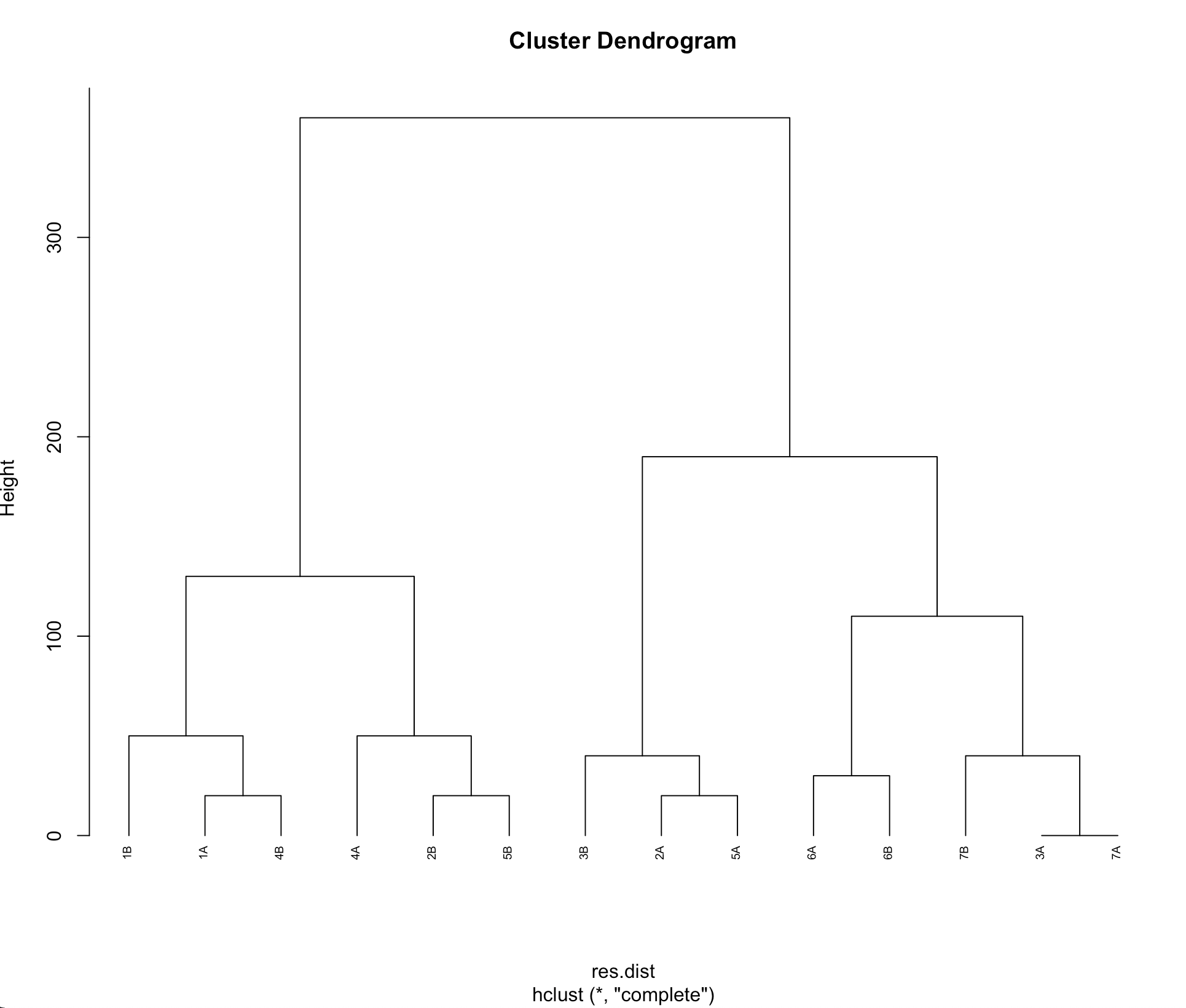
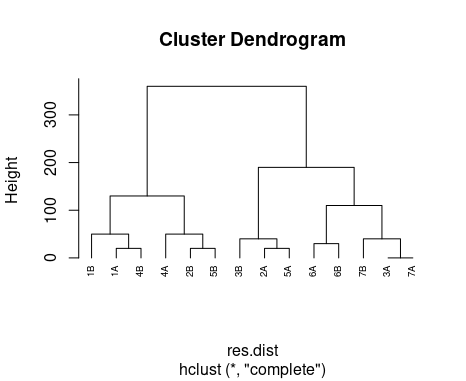
plot(x=hcl, hang = -1, cex = 0.6)
(sorry can't upload the picture for network issue) but after this in my picture intead 1A, 2A, 3A, 3B it comes 1,2,3,.....,14. how can i solve this?
CodePudding user response:
After dropping the 1st column, there is only one column left which collapses the data into a vector. Vector drops the rownames of the dataframe, hence there are no labels.
You can use drop = FALSE to keep the data as dataframe after the subset.
rownames(data) = data$Station
data = data[,-1, drop = FALSE]
res.dist = dist(x=data, method = "euclidean")
hcl = hclust(d=res.dist, method = "complete")
plot(x=hcl, hang = -1, cex = 0.6)
CodePudding user response:
There is no need to keep only the measurements column, the plot can be done by subsetting the data in the dist instruction.
Note that I use data[-1], not data[, -1], which would drop the dim attribute. The former always returns a sub-df.
rownames(data) <- data$Station
res.dist <- dist(x = data[-1], method = "euclidean")
hcl <- hclust(d = res.dist, method = "complete")
plot(x = hcl, hang = -1, cex = 0.6)