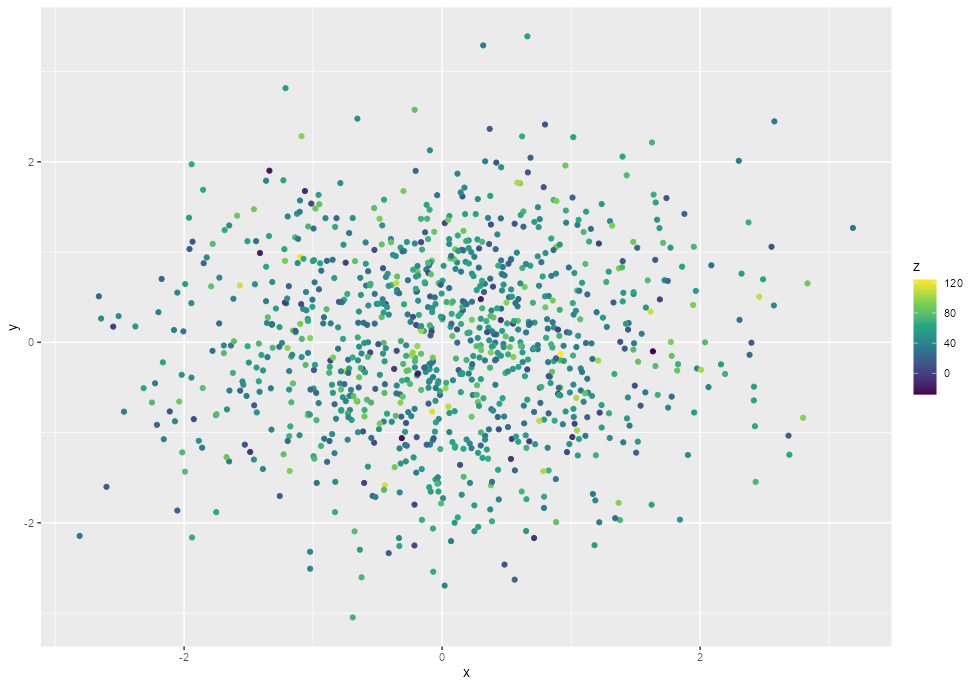
I often find myself working with data with long-tail distributions, so that a huge amount of range in values happens in the top 1-2% of the data. When I plot the data, the upper outliers cause variation in the rest of the data to wash out, but I want to show those difference.
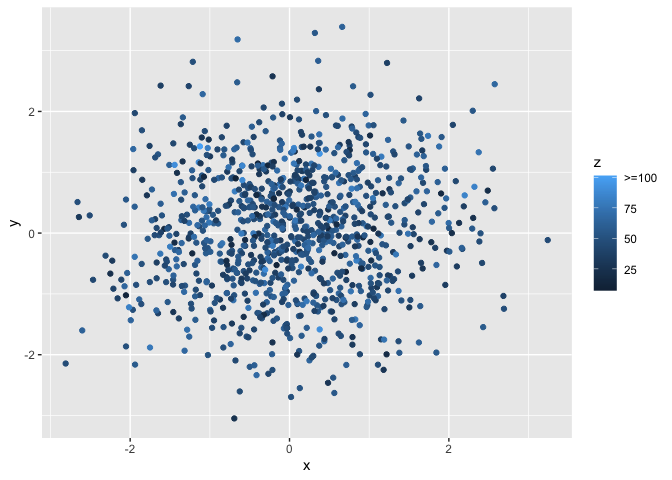
I know there are other ways of handling this, but I found that capping the values towards the end of the distribution and then applying a continuous color palette (i.e., in ggplot) is one way that works for me to represent the data. However, I want to ensure the legend stays accurate, by adding a >= sign to the last legend label
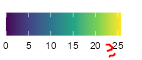
The picture below shows the of legend I'd like to achieve programmatically, with the >= sign drawn in messily in red.
I also know I can manually set breaks and labels, but I'd really like to just do something like, if(it's the last label) ~paste0(">=",label) else label) (to show with pseudo code)
Reproducible example: (I want to alter the plot legend to prefix just the last label)
set.seed(123)
x <- rnorm(1:1e3)
y <- rnorm(1:1e3)
z <- rnorm(1e3, mean = 50, sd = 15)
d <- tibble(x = x
,y = y
,z = z)
d %>%
ggplot(aes(x = x
,y = y
,fill = z
,color = z))
geom_point()
scale_color_viridis_c()
CodePudding user response:
One option would be to pass a function to the labels argument which replaces the last element or label with your desired label like so:
library(ggplot2)
set.seed(123)
x <- rnorm(1:1e3)
y <- rnorm(1:1e3)
z <- rnorm(1e3, mean = 50, sd = 15)
d <- data.frame(
x = x,
y = y,
z = z
)
ggplot(d, aes(
x = x,
y = y,
fill = z,
color = z
))
geom_point()
scale_fill_continuous(labels = function(x) {
x[length(x)] <- paste0(">=", x[length(x)])
x
}, aesthetics = c("color", "fill"))