I have some dataset similar to this:
df <- data.frame(n = seq(1:1000000), x = sample(LETTERS, 1000000, replace = T))
I'm looking for a guidance in finding a way to split variable x into multiple categorical variables with range 0-1
In the end it would look like this:
n x A B C D E F G H . . .
1 D 0 0 0 1 0 0 0 0 . . .
2 B 0 1 0 0 0 0 0 0 . . .
3 F 0 0 0 0 0 1 0 0 . . .
In my dataset, there's way more codes in variable x so adding each new variable manually would be too time consuming.
I was thinking about sorting codes in var x and assigning them an unique number each, then creating an iterating loop that creates new variable for each code in variable x. But i feel like i'm overcomplicating things
CodePudding user response:
Using match. First create a vector of zeroes, then match letter of df row with vector from the alphabet and turn to 1. You may use builtin LETTERS constant. Finally Vectorize the thing and cbind.
f <- \(x) {
z <- numeric(length(LETTERS))
z[match(x, LETTERS)] <- 1
setNames(z, LETTERS)
}
cbind(df, t(Vectorize(f)(df$x)))
# n x A B C D E F G H I J K L M N O P Q R S T U V W X Y Z
# Q 1 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
# E 2 E 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# A 3 A 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# Y 4 Y 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0
# J 5 J 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# D 6 D 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# R 7 R 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0
# Z 8 Z 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1
# Q.1 9 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
# O 10 O 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0
Alternatively, transform x to a factor with LETTERS as levels and use model.matrix.
df <- transform(df, x=factor(x, levels=LETTERS))
cbind(df, `colnames<-`(model.matrix(~ 0 x, df), LETTERS))
# n x A B C D E F G H I J K L M N O P Q R S T U V W X Y Z
# 1 1 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
# 2 2 E 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# 3 3 A 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# 4 4 Y 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0
# 5 5 J 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# 6 6 D 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
# 7 7 R 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0
# 8 8 Z 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1
# 9 9 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
# 10 10 O 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0
Data:
n <- 10
set.seed(42)
df <- data.frame(n = seq(1:n), x = sample(LETTERS, n, replace = T))
CodePudding user response:
A fast and easy way is to use fastDummies::dummy_cols:
fastDummies::dummy_cols(df, "x")
An alternative with tidyverse functions:
library(tidyverse)
df %>%
left_join(., df %>% mutate(value = 1) %>%
pivot_wider(names_from = x, values_from = value, values_fill = 0) %>%
relocate(n, sort(colnames(.)[-1])))
output
> dummmy <- fastDummies::dummy_cols(df, "x")
> colnames(dummy)[-c(1,2)] <- LETTERS
> dummy
n x A B C D E F G H I J K L M N O P Q R S T U V W X Y Z
1 1 Z 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1
2 2 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
3 3 E 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
4 4 H 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
5 5 T 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0
6 6 X 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0
7 7 R 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0
8 8 F 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
9 9 Z 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1
10 10 S 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0
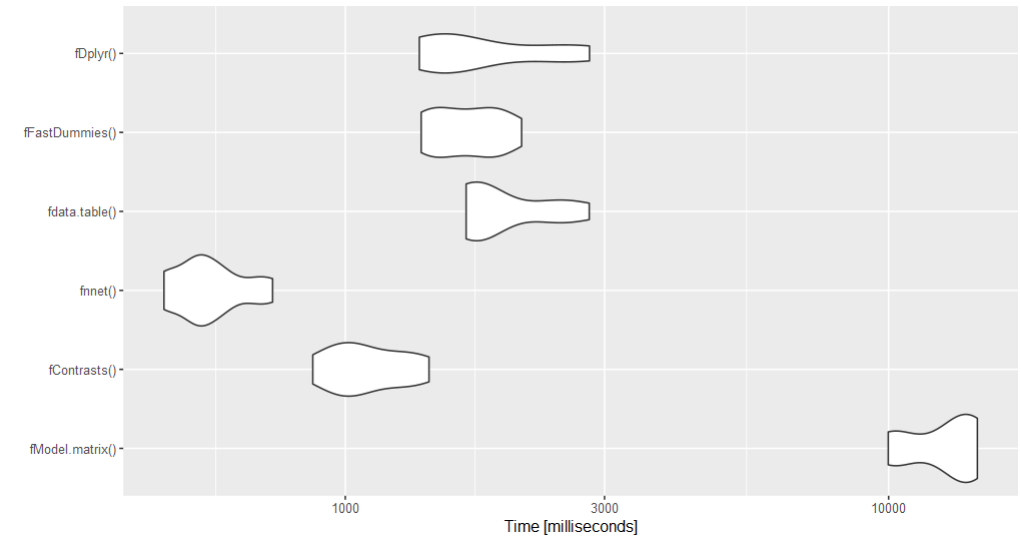
Benchmark Since there are many solutions and the question involves a large dataset, a benchmark might help. The nnet solution is the fastest according to the benchmark.
set.seed(1)
df <- data.frame(n = seq(1:1000000), x = sample(LETTERS, 1000000, replace = T))
library(microbenchmark)
bm <- microbenchmark(
fModel.matrix(),
fContrasts(),
fnnet(),
fdata.table(),
fFastDummies(),
fDplyr(),
times = 10L,
setup = gc(FALSE)
)
autoplot(bm)
CodePudding user response:
using data.table
library(data.table)
setDT(df) #make df a data.table if needed
merge(df, dcast(df, n ~ x, fun.agg = length), by = c("n"))
CodePudding user response:
The main question here is that of resources? I think. I found using nnet is a fast solution:
library(nnet)
library(dplyr)
df %>% cbind(class.ind(.$x) == 1) %>%
mutate(across(-c(n, x), ~.*1))
n x A B C D E F G H I J K L M N O P Q R S T U V W X Y Z
1 1 E 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
2 2 H 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
3 3 L 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0
4 4 M 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0
5 5 R 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0
6 6 A 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
7 7 Y 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0
8 8 Y 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0
9 9 F 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
10 10 U 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0
11 11 O 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0
12 12 I 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
13 13 O 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0
14 14 Z 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1
15 15 P 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0
16 16 T 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0
17 17 F 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
18 18 K 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
19 19 H 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
20 20 V 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0
21 21 V 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0
22 22 G 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
23 23 P 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0
24 24 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
25 25 V 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0
26 26 R 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0
27 27 Q 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0
28 28 B 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
29 29 D 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
30 30 M 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0
31 31 E 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
32 32 V 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0
33 33 S 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0
34 34 Y 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0
35 35 T 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0
[ reached 'max' / getOption("max.print") -- omitted 999965 rows ]
>
CodePudding user response:
You can use contrasts and match them to df$x. This will work also for more codes in variable x and not only for LETTERS.
. <- contrasts(as.factor(df$x), FALSE)
#. <- contrasts(as.factor(unique(df$x)), FALSE) #Alternative
cbind(df, .[match(df$x, rownames(.)),])
#cbind(df, .[fastmatch::fmatch(df$x, rownames(.)),]) #Alternative
# n x C I L T Y
#1 1 I 0 1 0 0 0
#2 2 C 1 0 0 0 0
#3 3 C 1 0 0 0 0
#4 4 Y 0 0 0 0 1
#5 5 L 0 0 1 0 0
#6 6 T 0 0 0 1 0
#...
Another option would be to use ==.
. <- unique(df$x)
cbind(df, do.call(cbind, lapply(setNames(., .), `==`, df$x)))
#cbind(df, sapply(unique(df$x), `==`, df$x)) #Alternative
Or indexing in a matrix.
. <- unique(df$x) #Could be sorted
#. <- collapse::funique(df$x) #Alternative
#. <- kit::funique(df$x) #Alternative
i <- match(df$x, .)
#i <- fastmatch::fmatch(df$x, .) #Alternative
#i <- data.table::chmatch(df$x, .) #Alternative
nc <- length(.)
nr <- length(i)
cbind(df, matrix(`[<-`(integer(nc * nr), 1:nr nr * (i - 1), 1), nr, nc,
dimnames=list(NULL, .)))
Or using outer.
. <- unique(df$x)
cbind(df, outer(df$x, setNames(., .), `==`))
Or using rep and m̀atrix`.
. <- unique(df$x)
n <- nrow(df)
cbind(df, matrix(df$x == rep(., each=n), n, dimnames=list(NULL, .)))