I have tens of thousands of images. I want to generate a histogram for each pixel. I have come up with the following code using NumPy to do this that works:
import numpy as np
import matplotlib.pyplot as plt
nimages = 1000
im_shape = (64,64)
nbins = 100
#predefine the histogram bins
hist_bins = np.linspace(0,1,nbins)
#create an array to store histograms for each pixel
perpix_hist = np.zeros((64,64,nbins))
for ni in range(nimages):
#create a simple image with normally distributed pixel values
im = np.random.normal(loc=0.5,scale=0.05,size=im_shape)
#sort each pixel into the predefined histogram
bins_for_this_image = np.searchsorted(hist_bins, im.ravel())
bins_for_this_image = bins_for_this_image.reshape(im_shape)
#this next part adds one to each of those bins
#but this is slow as it loops through each pixel
#how to vectorize?
for i in range(im_shape[0]):
for j in range(im_shape[1]):
perpix_hist[i,j,bins_for_this_image[i,j]] = 1
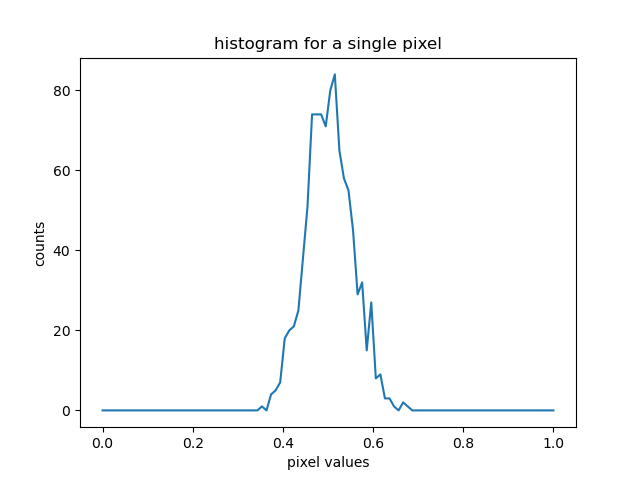
#plot histogram for a single pixel
plt.plot(hist_bins,perpix_hist[0,0])
plt.xlabel('pixel values')
plt.ylabel('counts')
plt.title('histogram for a single pixel')
plt.show()
I would like to know if anyone can help me vectorize the for loops? I can't think of how to index into the perpix_hist array properly. I have tens/hundreds of thousands of images and each image is ~1500x1500 pixels, and this is too slow.
CodePudding user response:
You can vectorize it using np.meshgrid and providing indices for first, second and third dimension (the last dimension you already have).
y_grid, x_grid = np.meshgrid(np.arange(64), np.arange(64))
for i in range(nimages):
#create a simple image with normally distributed pixel values
im = np.random.normal(loc=0.5,scale=0.05,size=im_shape)
#sort each pixel into the predefined histogram
bins_for_this_image = np.searchsorted(hist_bins, im.ravel())
bins_for_this_image = bins_for_this_image.reshape(im_shape)
perpix_hist[x_grid, y_grid, bins_for_this_image] = 1