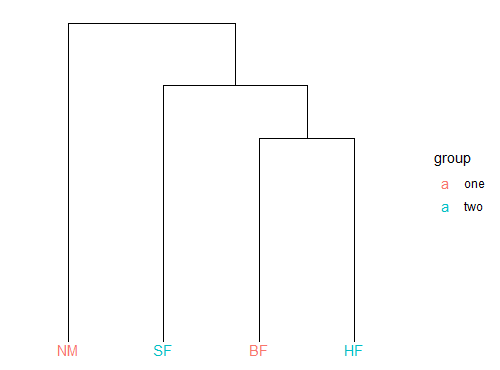Vegan::meandist() has a really nice plot method that creates a dendrogram of the mean dissimilarities. How can I incorporate the output into ggplot to have full control over the aesthetics? Here is some sample code using Dune. As an example, I'd like to recreate the dendrogram in ggplot and color each Management level by 'Use' (see factors in Dune.env).
# Species and environmental data
require(vegan)
dune <- read.delim ('https://raw.githubusercontent.com/zdealveindy/anadat-r/master/data/dune2.spe.txt', row.names = 1)
dune.env <- read.delim ('https://raw.githubusercontent.com/zdealveindy/anadat-r/master/data/dune2.env.txt', row.names = 1)
data(dune)
data(dune.env)
dune_dist <- vegdist(dune, method = "bray", na.rm=T)
dissim <- meandist(dune_dist, grouping = dune.env$Management)
plot(dissim)
CodePudding user response: